295 THE SHAPE OF PORCINE NEURAL PROGENITOR CELL CELLULAR GENEALOGIES REVEALED BY TIME-LAPSE IMAGING
I. Faerge A , A. Egeskov-Madsen A and P. Holm AInstitute of Basic Animal and Veterinary Sciences, Life Sciences, University of Copenhagen, Denmark
Reproduction, Fertility and Development 23(1) 245-245 https://doi.org/10.1071/RDv23n1Ab295
Published: 7 December 2010
Abstract
Porcine neural progenitor cells (pNPC) derived from embryonic stem cells are capable of self-renewal and differentiation into neural and glia lineages, rendering them promising candidates for cell-based therapy of neurodegenerative diseases in a large animal biomedical model. A prerequisite for the successful future therapeutic use of pNPC is a comprehensive characterisation and understanding of the neurogenic process. This is important for learning how to direct cell fates into required proportions of the cell type wanted for the specific brain disease to be treated, and it is crucial for avoiding uncontrolled cell proliferation leading to fatal tumour formations. Time-lapse analysis is a powerful tool to obtain live cell characterisation by analysing individual cell fate. Information on cellular development, division, and differentiation can be composed into a pedigree-like structure denoted as cellular genealogy giving an overview of the proliferation profile of a cell culture and the duration of each cell cycle (Al-Kofani et al. 2006). The aim of the study was to construct cellular genealogies of pNPC and differentiated neural lineages, respectively, by time-lapse imaging to evaluate the effect of external variables observed by changes in the topology of the cellular genealogy. Porcine NPC were derived from epiblast cells isolated from day-9 porcine blastocysts and cultured in DMEM/12, Pen/strep, B27 and N2 with basic fibroblast growth factor and epidermal growth factor, and differentiation was obtained by withdrawal of basic fibroblast growth factor and epidermal growth factor. The state of cellular development of undifferentiated and differentiated pNPC was verified immunohistochemically by the presence of SOX2, NESTIN, TUJI, and GFAB (Rasmussen et al. 2010). The time-lapse images were captured by a Nikon Biostation with a 10× resolution under phase contrast in a humidified chamber at 38°C with 5% CO2, 5% O2, and 90% N2. For each sequence, images were captured at intervals of 10 min in 16 frames. Sequences 1, 2, and 3 constituted passage 15 pNPC, passage 4 pNPC, and presumably differentiated cells, respectively. For each sequence, cell cycle length was calculated after manual tracking of selected cells. The cell cycle length of pNPC is shown in Table 1. Based on these data, cellular genealogies characteristic of each individual cell type have been constructed.
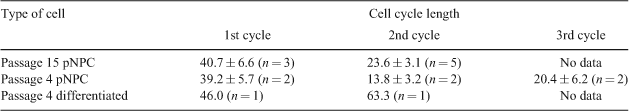
|


