Endo-siRNA deficiency results in oocyte maturation failure and apoptosis in porcine oocytes
Wanxin Liu A , Qi Zhao A , Shanhua Piao A , Chunsheng Wang A , Qingran Kong B C and Tiezhu An A CA Laboratory of Animal Developmental Biology, College of Life Science, Northeast Forestry University, Harbin, Heilongjiang Province 150040, China.
B Laboratory of Embryo Biotechnology, College of Life Science, Northeast Agricultural University, Harbin, Heilongjiang Province 150030, China.
C Corresponding authors. Emails: kqr721726@163.com; antiezhu@qq.com
Reproduction, Fertility and Development 29(11) 2168-2174 https://doi.org/10.1071/RD16498
Submitted: 29 August 2016 Accepted: 31 January 2017 Published: 12 April 2017
Journal Compilation © CSIRO 2017 Open Access CC BY-NC-ND
Abstract
Both microRNAs (miRNAs) and endogenous small interfering RNAs (endo-siRNAs) play key regulatory roles in gene expression. Some studies have demonstrated that the function of miRNA is suppressed in mouse oocytes, suggesting that endo-siRNA, not miRNA, is essential for female meiosis. This finding has yet to be confirmed in other species. In this study, by knockdown of DICER1, DROSHA and its cofactor DiGeorge syndrome critical region 8 (DGCR8) in porcine oocytes, we found that the proportion of oocytes with DICER1 deficiency that developed to meiosis II (MII) stage was significantly lower than oocytes with DROSHA and DGCR8 deficiency (39.23 versus 68.71 and 71.25% respectively; P < 0.05). Oocytes lacking DROSHA and DGCR8 formed a barrel-shaped metaphase I spindle, with chromosomes tightly aligned at the metaphase plate whereas most oocytes (87%) lacking DICER1 showed spindle abnormalities during oocyte in vitro maturation. Furthermore, DICER1 deficiency also resulted in oocyte apoptosis. These results indicate that endo-siRNAs are essential for oocyte maturation in pigs.
Additional keywords: gene regulation, in vitro maturation, livestock.
Introduction
Small RNAs are involved in RNA interference (RNAi) and include three classes: microRNAs (miRNAs), small interfering RNAs (siRNAs) and Piwi interacting RNAs (piRNAs; García-López et al. 2014). There are both endogenous siRNAs (endo-siRNAs) and exogenous siRNAs (exo-siRNAs). Gene expression can be artificially repressed by siRNAs (Saito and Siomi 2010; Claycomb 2014). PiRNAs (24–31 nucleotides (nt) in length) associated with Piwi-subfamily proteins are essential for male fertility (Ghildiyal and Zamore 2009; Kim et al. 2009; Reuter et al. 2011; Stein et al. 2015). The other two classes, miRNAs and endo-siRNAs, are both important for oocyte maturation. There are many similarities between miRNAs and endo-siRNAs including their length, both being about 21–23 nt (Hamilton et al. 2002; Ambros et al. 2003). All of these similarities make it hard to distinguish the function of endo-siRNAs from miRNAs. Therefore, the functions of miRNAs and endo-siRNAs need to be further clarified.
Dicer is involved in biogenesis of both miRNAs and endo-siRNAs while Drosha and DiGeorge syndrome critical region 8 (DGCR8) are only needed in miRNA biogenesis (Bernstein et al. 2001; Murchison et al. 2007; Kim et al. 2009). In previous studies, researchers studied small RNA function in Dicer knockout (KO) oocytes and zygotes and suggested the abnormalities were caused by a miRNA deficiency (Murchison et al. 2007; Tang et al. 2007). Recently, scientists knocked out Drosha and DGCR8 and neither of the KO oocytes was a phenocopy of the Dicer KOs, indicating that endo-siRNAs may play an important role (Ma et al. 2010; Suh et al. 2010; Yuan et al. 2014). In these studies of mouse, neither Drosha nor DGCR8 deficiency resulted in inaccurate oocyte maturation while Dicer deficiency caused oocyte maturation failure. Moreover, Argonaute (AGO) proteins play an indispensible role in target regulation of both endo-siRNAs and miRNAs. However, miRNAs bind to partially complementary sites in target mRNA 3′ untranslated regions (UTRS) and cause translational repression and mRNA decay through association with any of the four AGO proteins (AGO1–4), whereas endo-siRNAs bind to Argonaute 2 (AGO2), the only mammalian AGO protein thought to possess endonucleolytic activity, and mediate endonucleolytic cleavage of target mRNA (Watanabe et al. 2006; Ghildiyal and Zamore 2009). Endo-siRNAs regulate gene expression through their duplexes with AGO2. Only one of the two strands, the ‘guide’ strand, is incorporated into the multi-protein RNA-induced silencing complex (RISC); the other (‘passenger’) strand is discarded (Tam et al. 2008; Czech and Hannon 2011; Nejepinska et al. 2012). The guide strand recognises a target mRNA by Watson–Crick base pairing and based on the degree of sequence complementarity between the siRNAs and target mRNA, either endonucleolytic cleavage or translational repression of the target mRNA follows (Carthew and Sontheimer 2009). Thus, it is clear that AGO2 is necessary for endo-siRNA function. Inactivation of AGO2 led to the same results as Dicer inactivation, which further confirmed that endo-siRNAs were indispensible during oocyte maturation (Kaneda et al. 2009; Stein et al. 2015). However, the results have only been obtained in the mouse model, so further studies will be required to understand the function of endo-siRNAs in other species.
Pigs (Sus scrofa) are an important species for disease modelling, biomedical research and food production. Pigs are important not only in agriculture but also in biomedicine. In the field of biomedicine, pigs are more anatomically and physiologically analogous to humans than mice (Pratt et al. 2006; Whyte and Prather 2011). Alterations of porcine key genes in the reproductive pathway provide model animals to improve our understanding of the causes and potential treatments of many human reproductive disorders. Therefore, in this study, we examined the functions of miRNAs and endo-siRNAs in porcine oocytes by knockdown of oocyte DICER1, DROSHA and DGCR8. We found that the absence of DICER1, but not of DROSHA or DGCR8, resulted in spindle abnormalities and oocyte apoptosis during oocyte in vitro maturation in pig, indicating that endo-siRNA rather than miRNA is essential for oocyte maturation.
Materials and methods
Oocyte collection and in vitro maturation (IVM)
Porcine ovaries were collected from a local slaughter house and kept in 0.9% saline with antibiotics at 37°C. Antral follicles whose diameter was between 3 and 5 mm were aspirated with an appropriate needle. Aspirated oocytes with an evenly granulated cytoplasm and at least three uniform layers of compact cumulus cells were selected and washed three times with maturation medium (TCM199 (Invitrogen) plus 0.05 mg mL−1 epidermal growth factor, 0.5 mg mL−1 LH and 0.5 mg mL−1 FSH (all Sigma-Aldrich)). The oocytes with or without microinjection were cultured in 24-well plates (Corning) containing 500 mL of maturation medium at 39°Cin 5% CO2 in air and saturated humidity (Kong et al. 2014).
Microinjection
To perform DICER1, DROSHA and DGCR8 knockdown experiments, the granulosa cells of oocytes at germinal vesicle (GV) stage were denuded. Then locked nucleic acid (LNA)-siRNA microinjections were carried out with an Eppendorf FemtoJet microinjector and Narishige NT-88NE micromanipulators. For injection, a glass capillary Femtotip II (Eppendorf) was loaded with 10 pL of 10 μM LNA-DICER1, LNA-DROSHA or LNA-DGCR8 (Exiqon) by microloader (Eppendorf) and the solution was injected into the cytoplasm of GV oocytes in a 200-μL drop of manipulation medium (TCM-199 (Invitrogen) plus 30 mg mL−1 bovine serum albumin (BSA)) supplemented with 7.5 μg mL−1 cytochalasin B. The injection conditions consisted of 250 hPa injection pressure, 60 hPa comensation pressure and 0.7 s injection time. Immediately after microinjection, oocytes were washed and co-cultured with mural granulosa cells in maturation medium. A scrambled LNA-siRNA (Exiqon) was used as a negative control (NC; Wei et al. 2011).
Quantitative real-time polymerase chain reaction (PCR)
Total RNA was extracted using the PureLink Micro-to-Midi System (Invitrogen) according to the manufacturer’s instructions and reverse transcription was used to generate cDNAs using the PrimeScript RT Reagent kit (TaKaRa). Real-time PCR was performed using SYBR Premix Ex Taq (TaKaRa) and the 7500 Real-Time PCR System (Applied Biosystems). The reaction parameters were 95°C for 30 s followed by 40 two-step cycles of 95°C for 5 s and 60°C for 34 s. All the primer pairs used for PCR amplification are shown in Table S1, available as Supplementary Material to this paper. Ct values were calculated using Sequence Detection System software (Applied Biosystems) and the amount of target sequence normalised to the reference sequence was calculated by the ΔΔCT method. The 18SrRNA was used as control. The oocytes without injection were used as a reference sample. Each pool contained 50 oocytes. All data are based on biological triplicates.
Western blot
Proteins from 100 oocytes at the appropriate stage of maturation were collected in sodium dodecyl sulfate (SDS) sample buffer and heated for 10 min at 100°C. After cooling on ice and centrifugation at 10 000g for 3 min at room temperature, samples were frozen at −80°C until use. The total proteins were separated by polyacrylamide gel electrophoresis with SDS (SDS-PAGE) with a 4% stacking gel and a 9% separating gel at 90 V, 0.5 h and 110 V, 2.5 h respectively and were then electrophoretically transferred onto a polyvinylidene difluoride membrane for 1 h, 350 mA at 4°C. Membranes were blocked in phosphate-buffered saline-Tris (PBST) buffer (10 mM Tris, 150 mM NaCl, 0.1% Tween 20, pH 7.4) containing 5% BSA (blocking solution) for 1 h at room temperature and then incubated with antibody against DICER1 (ab14601; Abcam) or p53 (sc-65226; Santa Cruz Biotechnology) diluted 1 : 2000 in blocking solution overnight at 4°C. After three 10-min washes in PBST, the membrane was incubated with a secondary antibody against mouse IgG (A9044; Sigma) diluted 1 : 10 000 in PBST for 1 h at 37°C. After being washed three times for 10 min each, the membrane was processed using the enhanced chemiluminescence (ECL) detection system (Amersham). Equal protein loading was confirmed by detection of β-actin (A1978; Sigma; Kong et al. 2014).
Immunofluorescence and confocal microscopy
After removing the zona pellucida in acidic Tyrode’s solution (pH 2.5), oocytes were fixed with 4% paraformaldehyde in PBS (pH 7.4) for at least 30 min at room temperature. Oocytes were permeabilised with 1% Triton X-100 overnight at 37°C, followed by blocking in PBS containing 1% BSA (blocking solution) for 1 h and incubation overnight at 4°C with antibody against β-tubulin (T5201; Sigma) diluted 1 : 500 in blocking solution. After three washes in PBS containing 0.1% Tween 20 and 0.01% Triton X-100 (washing solution) for 5 min each, the oocytes were labelled with antibody against alpha-fetoprotein (AFP) (H00000174-M01; Abnova) diluted 1 : 500 in washing solution for 1 h at room temperature. After one wash in washing solution, the nuclear status of the oocytes was evaluated by staining with Hoechst 33342 (5 μg mL−1 in washing solution; Sigma-Aldrich) for 2 min. After another three washes in washing solution for 8 min each, oocytes were mounted on glass slides in ProLong Diamond Antifade Mountant reagent (Life Technologies). Cells were observed under a confocal laser-scanning microscope (Leica TCS SP2 AOBS) as soon as possible after preparation. Each experiment was repeated three times and at least 30 oocytes were examined each time. In addition, the same instrument settings were used for each replicate (Xu et al. 2009; Kong et al. 2014).
Annexin-V assay
An annexin-V conjugate (Molecular Probes) was used to identify phosphatidylserine exteriorisation in apoptotic cells. According to the manufacturer’s instructions, oocytes without zonae pellucidae were washed twice in PBS at 4°C and then washed three times in binding buffer. They were then incubated with annexin-V for 15 min before being transferred to a 0.25 mg mL−1 solution of propidium iodide (Sigma-Aldrich) to allow recognition of necrotic cells. Oocytes were washed three times in binding buffer before mounting in ProLong Diamond Antifade Mountant reagent (Life Technologies) and analysing via fluorescence microscopy.
Statistical analysis
Statistical analysis was performed using SPSS 13.0 for MicroSoft Windows (IBM). Data are shown as the mean ± s.e.m. One-way analysis of variance was used to assess any differences between groups. The Duncan method was used for pairwise comparisons followed by a Bonferroni correction. P < 0.05 (two-tailed) was considered to be statistically significant.
Results
Efficient knockdown of DICER1, DROSHA and DGCR8
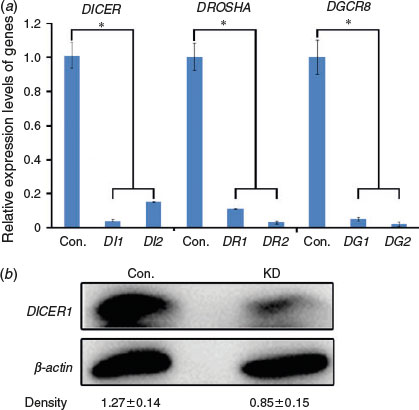
To obtain DICER1, DROSHA and DGCR8 knockdown porcine oocytes, we synthesised two locked nucleic acids (LNAs) for each gene and microinjected the LNAs into oocytes at the GV stage. Quantitative PCR was performed after 24 h of microinjection to test the knockdown efficiencies; efficient knockdown of DICER1, DROSHA and DGCR8 in porcine oocytes was confirmed (Fig. 1a). Moreover, a dramatic decrease in DICER1 protein was observed by western blot analysis (Fig. 1b). Therefore, we were successful in knockdown of DICER1, DROSHA and DGCR8 in porcine oocytes.
|
Knockdown of DICER1 results in nuclear maturation failure and abnormal chromosome alignment
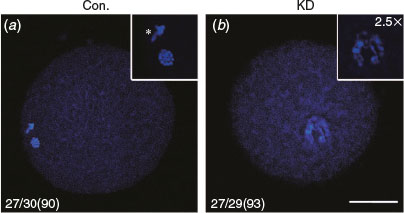
We examined the effect of DICER1, DROSHA and DGCR8 knockdown on nuclear maturation of porcine oocytes by LNA microinjection. The results showed that the rates of first polar body (PB1) extrusion showed no significant difference between control, scrambled (negative control) LNA, DROSHA-LNA and DGCR8-LNA groups (75.2 and 69.75 versus 68.71 and 71.25% respectively; P > 0.05; Table 1), but the proportion of oocytes that developed to metaphase II (MII) stage in the DICER1-LNA group was significantly lower than in the control and scrambled LNA groups (39.23 versus 75.2 and 69.75% respectively; P < 0.05; Table 1). Considering that DICER knockdown oocytes cannot extrude the PB1, we decided to observe the nuclear morphology by Hochest33342 staining. The PB1 and well-organised chromosomes were found in the oocytes without injection (n = 27; 90%; Fig. 2a) whereas the DICER1 knockdown oocytes displayed unaligned chromosomes (n = 27; 93%; Fig. 2b). Given that DICER1 is important for endo-siRNA biogenesis while DROSHA and DGCR8 are dispensable, we believed that endo-siRNA not miRNA deficiency led to the abnormal PB1 extrusion and chromosome alignment.

|
|
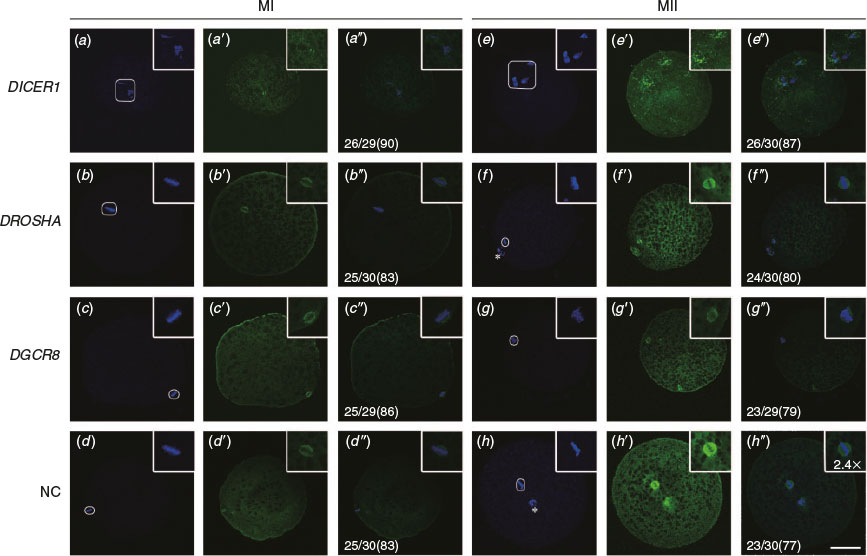
Oocyte DICER1 knockdown leads to disordered spindle morphology
Earlier studies showed that Dicer rather than Drosha or DGCR8 knockdown can regulate the formation of the spindle in mouse oocytes. In the study, immunofluorescence analysis of the spindle using β-tubulin antibody was performed to determine the progression of meiosis in porcine oocytes. Results revealed that 83%, 80%, 73% and 93% of oocytes arrived at the metaphase I (MI), anaphase I (AI), telophase I (TI) and MII stages at 18 h, 20 h, 22 h and 42 h during IVM respectively (n = 25, 24, 22 and 28 respectively; see Fig. S1, available as Supplementary Material to this paper). We found that oocytes from the DROSHA-LNA, DGCR8-LNA and scrambled LNA groups proceeded to form a barrel-shaped metaphase I spindle, with chromosomes tightly aligned at the metaphase plate (Fig. 3b–b″, c–c″, d–d″), and, by the extrusion of PB1, completed meiosis I and arrested at the metaphase stage of meiosis II (Fig. 3f–f″, g–g″, h–h″). In contrast, in most oocytes of the DICER1-LNA group, the chromosomes remained dispersed and never aligned and the spindle was extraordinarily disorganised (Fig. 3a–a″, e–e″). Considering that DICER1 is indispensible for endo-siRNAs while DROSHA and DGCR8 are not, these results suggest that endo-siRNA is indispensible for chromosome alignment and spindle formation of porcine oocytes. In other words, endo-siRNA is necessary for meiosis of porcine oocytes.
Oocyte DICER1 deficiency results in apoptosis
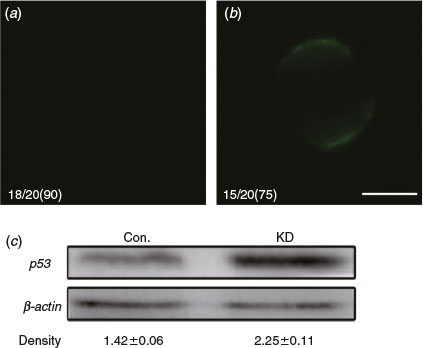
We have shown that the absence of DICER1 leads to immature oocytes in pigs. However, it is unknown whether the knockout of DICER1 causes oocyte apoptosis. To detect phosphatidylserine (PS) exteriorisation (a signal of early apoptosis) in MII-stage oocytes, we use fluorescent annexin-V that can only combine with PS outside of the membrane with Ca2+ (Andree et al. 1990; van Engeland et al. 1998). Results showed 75% of oocytes in the DICER1-LNA group were annexin-V-positive (n = 15; Fig. 4b) whereas annexin-V-positive oocytes were significantly fewer in the control group (10%; n = 2; Fig. 4a). Further, western blotting of MII-stage oocytes showed that the level of p53 protein in the DICER1-LNA group was dramatically higher than in the control group (Fig. 4c). Together, these results indicate that endo-siRNA deficiency leads to oocyte apoptosis.
|
Discussion
In this study, the lack of DICER1 but not of DROSHA or DGCR8 resulted in spindle abnormalities and oocyte apoptosis during oocyte in vitro maturation in the pig, indicating that endo-siRNA rather than miRNA is indispensible for oocyte maturation.
It has been reported that a lack of mouse endo-siRNAs results in some female reproductive diseases such as premature ovarian failure and infertility (Yuan et al. 2014). However, the phenomenon was not demonstrated in other species. In this study, we examined whether porcine endo-siRNA deficiency resulted in any defects like in the mouse. We knocked down porcine oocyte DICER1, DROSHA and DGCR8 by microinjection of their LNAs which can knockdown genes efficiently (Kanwar et al. 2015; Jolly et al. 2016). By quantitative real-time PCR we confirmed that the LNAs for these three genes were effective. Considering that DICER1 knockdown caused oocyte maturation failure (rather than DROSHA or DGCR8 as in the mouse), we further checked the porcine DICER1 knockdown effect by western blot (Fig. 1). So far, we believe that this is the first time that DICER1, DROSHA and DGCR8 have been knocked down in porcine oocytes.
The extrusion of the first polar body is an important mark for oocyte maturation (Evans and Robinson 2011; Schmerler and Wessel 2011). It has been found that Dicer deficiency causes improper oocyte maturation due to failure of PB1 extrusion in the mouse (Ma et al. 2010; Suh et al. 2010). We knocked down DICER1, DROSHA and DGCR8 in porcine oocytes and found that DICER1 deficiency caused PB1 extrusion failure whereas DROSHA and DGCR8 deficiency did not. To ensure this phenotype was a result of DICER deficiency rather than microinjection, we microinjected the scrambled LNA into oocytes and found normal PB1 extrusion (Table 1). Furthermore, the staining results showed that DICER1-deficient oocytes exhibited abnormal chromosome morphology and did not complete meiosis I (Fig. 2). Some studies demonstrated that Dicer deficiency led to spindle formation failure in mouse oocytes, which could be a cause of chromosome disorder. Thus, we performed immunofluorescence to check the chromosome and spindle morphology. The DROSHA and DGCR8 knockdown oocytes showed exactly the same chromosome and spindle morphology as both control and negative control groups (Fig. 3). It is believed that Dicer is involved in both endo-siRNA and miRNA biogenesis whereas Drosha and DGCR8 only function in miRNA biogenesis, thus we concluded that a lack of endo-siRNA and not miRNA was contributing to the abnormalities exhibited. Why does endo-siRNA deficiency cause disordered chromosomes and spindle morphology? The explanation could be that centromeres are composed of repetitive sequences and endo-siRNAs are involved in silencing of repetitive sequences (Bagasra and Prilliman 2004; Ekwall 2007; Banisch et al. 2012; Fukagawa and Earnshaw 2014). The loss of endo-siRNAs may activate heterochromatin repetitive sequences and prevent centromeres from connecting to microtubules (Kanellopoulou et al. 2005; Ekwall 2007; Saito and Siomi 2010; Castel and Martienssen 2013; Li 2014). As a result, the spindle cannot form properly during oocyte maturation.
It has been shown in some studies that abnormal meiosis causes oocyte apoptosis (Ene et al. 2013; Tripathi and Chaube 2015). Therefore, we examined apoptosis in the DICER1-deficient oocytes. Under knockdown of DICER1, we showed apoptosis in porcine oocytes by testing PS exteriorisation and p53 expression (Fig. 4), suggesting that endo-siRNA deficiency leads to oocyte apoptosis. Endo-siRNAs are so important in RISC that they are involved in regulation of gene expression. As a result, endo-siRNA deficiency may lead to abnormal expression of the gene network during oocyte maturation and result in oocyte apoptosis (Hussein et al. 2006; Coticchio et al. 2015).
Our findings indicate that the function of endo-siRNAs in porcine oocyte maturation is more significant than in the mouse. Pigs are more anatomically and physiologically analogous to humans than are mice, so further study of the reproductive diseases in humans is aided by studies in pigs. In summary, our findings provide two major insights into the roles of small RNAs in porcine oocytes by DICER1 knockdown. First, endo-siRNAs instead of miRNAs are essential for nuclear maturation of porcine oocytes. Second, the loss of endo-siRNAs leads to oocyte apoptosis.
* These authors contributed equally to this work.
Acknowledgement
This work was supported by the National Natural Science Foundation of China (grant number 31470079) and Academic Backbone Project of North-East Agricultural University (15XG19).
References
Ambros, V., Lee, R. C., Lavanway, A., Williams, P. T., and Jewell, D. (2003). MicroRNAs and other tiny endogenous RNAs in C. elegans. Curr. Biol. 13, 807–818.| MicroRNAs and other tiny endogenous RNAs in C. elegans.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD3sXjvFamsbs%3D&md5=96bf56bbce987562ee4ca55a18b90e9aCAS |
Andree, H. A., Reutelingsperger, C. P., Hauptmann, R., Hemker, H. C., Hermens, W. T., and Willems, G. M. (1990). Binding of vascular anticoagulant alpha (VAC alpha) to planar phospholipid bilayers. J. Biol. Chem. 265, 4923–4928.
| 1:CAS:528:DyaK3cXhvFCjtbw%3D&md5=67f3b944f935048c5779b7bf11bc0333CAS |
Bagasra, O., and Prilliman, K. R. (2004). RNA interference: the molecular immune system. J. Mol. Histol. 35, 545–553.
| 1:CAS:528:DC%2BD2cXhtFWqsbvJ&md5=bb83b1dd7f6fdef45aacc3e88b91dc93CAS |
Banisch, T. U., Goudarzi, M., and Raz, E. (2012). Small RNAs in germ cell development. Curr. Top. Dev. Biol. 99, 79–113.
| Small RNAs in germ cell development.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC38XmslClur0%3D&md5=c0299cc00ce1d5d2b45c36eb2461ae59CAS |
Bernstein, E., Caudy, A. A., Hammond, S. M., and Hannon, G. J. (2001). Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366.
| Role for a bidentate ribonuclease in the initiation step of RNA interference.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD3MXms12ksA%3D%3D&md5=9aa7ce0f3f150af438e925b4602badd8CAS |
Carthew, R. W., and Sontheimer, E. J. (2009). Origins and mechanisms of miRNAs and siRNAs. Cell 136, 642–655.
| Origins and mechanisms of miRNAs and siRNAs.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD1MXkvFGksb0%3D&md5=928572bba7e96f65e21366f1f8e71465CAS |
Castel, S. E., and Martienssen, R. A. (2013). RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat. Rev. Genet. 14, 100–112.
| RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3sXptlOrtQ%3D%3D&md5=a3c26290a9a4ca12765bb3092dca6aa0CAS |
Claycomb, J. M. (2014). Ancient endo-siRNA pathways reveal new tricks. Curr. Biol. 24, R703–R715.
| Ancient endo-siRNA pathways reveal new tricks.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC2cXhtlSqtrrJ&md5=d0910e31e394a11b0ea8557569efeaf9CAS |
Coticchio, G., Dal Canto, M., Mignini Renzini, M., Guglielmo, M. C., Brambillasca, F., Turchi, D., Novara, P. V., and Fadini, R. (2015). Oocyte maturation: gamete–somatic cell interactions, meiotic resumption, cytoskeletal dynamics and cytoplasmic reorganization. Hum. Reprod. Update 21, 427–454.
| Oocyte maturation: gamete–somatic cell interactions, meiotic resumption, cytoskeletal dynamics and cytoplasmic reorganization.Crossref | GoogleScholarGoogle Scholar |
Czech, B., and Hannon, G. J. (2011). Small RNA sorting: matchmaking for Argonautes. Nat. Rev. Genet. 12, 19–31.
| Small RNA sorting: matchmaking for Argonautes.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3cXhsFOiu7rE&md5=88be7b84a756225a01ed2319b77338d8CAS |
Ekwall, K. (2007). Epigenetic control of centromere behavior. Annu. Rev. Genet. 41, 63–81.
| Epigenetic control of centromere behavior.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD1cXns1Sjsg%3D%3D&md5=23d2f8d35abb41684c29ed41385758a7CAS |
Ene, A. C., Park, S., Edelmann, W., and Taketo, T. (2013). Caspase 9 is constitutively activated in mouse oocytes and plays a key role in oocyte elimination during meiotic prophase progression. Dev. Biol. 377, 213–223.
| Caspase 9 is constitutively activated in mouse oocytes and plays a key role in oocyte elimination during meiotic prophase progression.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3sXjtl2jsLg%3D&md5=17080a649bf510fa5d2925c708092ab0CAS |
Evans, J. P., and Robinson, D. N. (2011). The spatial and mechanical challenges of female meiosis. Mol. Reprod. Dev. 78, 769–777.
| The spatial and mechanical challenges of female meiosis.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3MXhtlagu7vF&md5=98324720abb82cb9614b48477f443c2eCAS |
Fukagawa, T., and Earnshaw, W. C. (2014). The centromere: chromatin foundation for the kinetochore machinery. Dev. Cell 30, 496–508.
| The centromere: chromatin foundation for the kinetochore machinery.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC2cXhsFagsbfF&md5=72ccdf824ecad0a65d5acc8622803d7aCAS |
García-López, J., Hourcade Jde, D., Alonso, L., Cardenas, D. B., and del Mazo, J. (2014). Global characterization and target identification of piRNAs and endo-siRNAs in mouse gametes and zygotes. Biochim. Biophys. Acta 1839, 463–475.
| Global characterization and target identification of piRNAs and endo-siRNAs in mouse gametes and zygotes.Crossref | GoogleScholarGoogle Scholar |
Ghildiyal, M., and Zamore, P. D. (2009). Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 10, 94–108.
| Small silencing RNAs: an expanding universe.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD1MXlvFChtA%3D%3D&md5=ec8c290c42b6f7fb65dff44f1be15800CAS |
Hamilton, A., Voinnet, O., Chappell, L., and Baulcombe, D. (2002). Two classes of short interfering RNA in RNA silencing. EMBO J. 21, 4671–4679.
| Two classes of short interfering RNA in RNA silencing.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD38Xms1KmsrY%3D&md5=edba38d4bcefe9ad2983f46594306fc5CAS |
Hussein, T. S., Thompson, J. G., and Gilchrist, R. B. (2006). Oocyte-secreted factors enhance oocyte developmental competence. Dev. Biol. 296, 514–521.
| Oocyte-secreted factors enhance oocyte developmental competence.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD28XotV2gsb4%3D&md5=02562f698c762d378021f5cb17dd2107CAS |
Jolly, P., Estrela, P., and Ladomery, M. (2016). Oligonucleotide-based systems: DNA, microRNAs, DNA/RNA aptamers. Essays Biochem. 60, 27–35.
| Oligonucleotide-based systems: DNA, microRNAs, DNA/RNA aptamers.Crossref | GoogleScholarGoogle Scholar |
Kaneda, M., Tang, F., O’Carroll, D., Lao, K., and Surani, M. A. (2009). Essential role for Argonaute2 protein in mouse oogenesis. Epigenetics Chromatin 2, 9.
| Essential role for Argonaute2 protein in mouse oogenesis.Crossref | GoogleScholarGoogle Scholar |
Kanellopoulou, C., Muljo, S. A., Kung, A. L., Ganesan, S., Drapkin, R., Jenuwein, T., Livingston, D. M., and Rajewsky, K. (2005). Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 19, 489–501.
| Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD2MXhs1OhsL4%3D&md5=efceee864c451a5a56a2981e78dc8c75CAS |
Kanwar, J. R., Roy, K., Maremanda, N. G., Subramanian, K., Veedu, R. N., Bawa, R., and Kanwar, R. K. (2015). Nucleic acid-based aptamers: applications, development and clinical trials. Curr. Med. Chem. 22, 2539–2557.
| Nucleic acid-based aptamers: applications, development and clinical trials.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC2MXhsFWms7%2FN&md5=d13a84fe285c88563e829cb3c160d670CAS |
Kim, V. N., Han, J., and Siomi, M. C. (2009). Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139.
| Biogenesis of small RNAs in animals.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD1MXpsVCitA%3D%3D&md5=07527daad2b4c416a50d9c8f0719380bCAS |
Kong, Q., Xie, B., Li, J., Huan, Y., Huang, T., Wei, R., Lv, J., Liu, S., and Liu, Z. (2014). Identification and characterization of an oocyte factor required for porcine nuclear reprogramming. J. Biol. Chem. 289, 6960–6968.
| Identification and characterization of an oocyte factor required for porcine nuclear reprogramming.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC2cXjvV2jur8%3D&md5=7cf5a1e90dc594e3033c010bfe0107feCAS |
Li, L. C. (2014). Chromatin remodeling by the small RNA machinery in mammalian cells. Epigenetics 9, 45–52.
| Chromatin remodeling by the small RNA machinery in mammalian cells.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC2cXhtVyiurvM&md5=f3036118ea960b10de4bc11ec6d54c72CAS |
Ma, J., Flemr, M., Stein, P., Berninger, P., Malik, R., Zavolan, M., Svoboda, P., and Schultz, R. M. (2010). MicroRNA activity is suppressed in mouse oocytes. Curr. Biol. 20, 265–270.
| MicroRNA activity is suppressed in mouse oocytes.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3cXhslSmurY%3D&md5=70bc0bb8228680522216090e03880864CAS |
Murchison, E. P., Stein, P., Xuan, Z., Pan, H., Zhang, M. Q., Schultz, R. M., and Hannon, G. J. (2007). Critical roles for Dicer in the female germline. Genes Dev. 21, 682–693.
| Critical roles for Dicer in the female germline.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD2sXjs1WqsbY%3D&md5=ec0372ecbcf37f287602c5f47ff61073CAS |
Nejepinska, J., Malik, R., Filkowski, J., Flemr, M., Filipowicz, W., and Svoboda, P. (2012). dsRNA expression in the mouse elicits RNAi in oocytes and low adenosine deamination in somatic cells. Nucleic Acids Res. 40, 399–413.
| dsRNA expression in the mouse elicits RNAi in oocytes and low adenosine deamination in somatic cells.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC38Xit1Ggsg%3D%3D&md5=fd0174bc6c1aa7eb85e4a0bb981b3e95CAS |
Pratt, S. L., Sherrer, E. S., Reeves, D. E., and Stice, S. L. (2006). Factors influencing the commercialisation of cloning in the pork industry. Soc. Reprod. Fertil. Suppl. 62, 303–315.
| 1:STN:280:DC%2BD28vktlamsA%3D%3D&md5=c501a5031d50b4086cc4afa2342689abCAS |
Reuter, M., Berninger, P., Chuma, S., Shah, H., Hosokawa, M., Funaya, C., Antony, C., Sachidanandam, R., and Pillai, R. S. (2011). Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480, 264–267.
| Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3MXhsFWqtb7P&md5=1a36f171104637d354f1a18626868eecCAS |
Saito, K., and Siomi, M. C. (2010). Small RNA-mediated quiescence of transposable elements in animals. Dev. Cell 19, 687–697.
| Small RNA-mediated quiescence of transposable elements in animals.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3cXhsValsLnM&md5=c77f154215847e9658aa81b9bc772f69CAS |
Schmerler, S., and Wessel, G. M. (2011). Polar bodies – more a lack of understanding than a lack of respect. Mol. Reprod. Dev. 78, 3–8.
| Polar bodies – more a lack of understanding than a lack of respect.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3MXpvV2lug%3D%3D&md5=94cff0daf98d766d1f0ec0a8baf57ec8CAS |
Stein, P., Rozhkov, N. V., Li, F., Cardenas, F. L., Davydenko, O., Vandivier, L. E., Gregory, B. D., Hannon, G. J., and Schultz, R. M. (2015). Essential role for endogenous siRNAs during meiosis in mouse oocytes. PLoS Genet. 11, e1005013.
| Essential role for endogenous siRNAs during meiosis in mouse oocytes.Crossref | GoogleScholarGoogle Scholar |
Suh, N., Baehner, L., Moltzahn, F., Melton, C., Shenoy, A., Chen, J., and Blelloch, R. (2010). MicroRNA function is globally suppressed in mouse oocytes and early embryos. Curr. Biol. 20, 271–277.
| MicroRNA function is globally suppressed in mouse oocytes and early embryos.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3cXhslSmurc%3D&md5=4843abb563228780190d10aa72ddacfaCAS |
Tam, O. H., Aravin, A. A., Stein, P., Girard, A., Murchison, E. P., Cheloufi, S., Hodges, E., Anger, M., Sachidanandam, R., Schultz, R. M., and Hannon, G. J. (2008). Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453, 534–538.
| Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD1cXmt1OrtLg%3D&md5=4ae7a8d452793f79982827a4b37e760dCAS |
Tang, F., Kaneda, M., O’Carroll, D., Hajkova, P., Barton, S. C., Sun, Y. A., Lee, C., Tarakhovsky, A., Lao, K., and Surani, M. A. (2007). Maternal microRNAs are essential for mouse zygotic development. Genes Dev. 21, 644–648.
| Maternal microRNAs are essential for mouse zygotic development.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD2sXjs1Wqsbo%3D&md5=e57024560755971541c04da28becd6bfCAS |
Tripathi, A., and Chaube, S. K. (2015). Roscovitine inhibits extrusion of second polar body and induces apoptosis in rat eggs cultured in vitro. Pharmacol. Rep. 67, 866–874.
| Roscovitine inhibits extrusion of second polar body and induces apoptosis in rat eggs cultured in vitro.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC28Xitlyjtb8%3D&md5=3bdaf05149317ae458c9283fe881dbc3CAS |
van Engeland, M., Nieland, L. J., Ramaekers, F. C., Schutte, B., and Reutelingsperger, C. P. (1998). Annexin V-affinity assay: a review on an apoptosis detection system based on phosphatidylserine exposure. Cytometry 31, 1–9.
| Annexin V-affinity assay: a review on an apoptosis detection system based on phosphatidylserine exposure.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DyaK1cXmtVGruw%3D%3D&md5=06db8efe35053c6b6ab2066289719becCAS |
Watanabe, T., Takeda, A., Tsukiyama, T., Mise, K., Okuno, T., Sasaki, H., Minami, N., and Imai, H. (2006). Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 20, 1732–1743.
| Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD28XmvV2ksLs%3D&md5=6b1683f26919055c0fe643d5e64eb881CAS |
Wei, Y., Huan, Y., Shi, Y., Liu, Z., Bou, G., Luo, Y., Zhang, L., Yang, C., Kong, Q., Tian, J., Xia, P., Sun, Q. Y., and Liu, Z. (2011). Unfaithful maintenance of methylation imprints due to loss of maternal nuclear Dnmt1 during somatic cell nuclear transfer. PLoS One 6, e20154.
| Unfaithful maintenance of methylation imprints due to loss of maternal nuclear Dnmt1 during somatic cell nuclear transfer.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3MXms1Kmurg%3D&md5=78c26b5b436f33ab0217fdc7ccfef241CAS |
Whyte, J. J., and Prather, R. S. (2011). Genetic modifications of pigs for medicine and agriculture. Mol. Reprod. Dev. 78, 879–891.
| Genetic modifications of pigs for medicine and agriculture.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BC3MXhtlagu7rK&md5=5f707bd4ee0269ee9e232a17fda0cc4bCAS |
Xu, B. Z., Li, M., Xiong, B., Lin, S. L., Zhu, J. Q., Hou, Y., Chen, D. Y., and Sun, Q. Y. (2009). Involvement of calcium/calmodulin-dependent protein kinase kinase in meiotic maturation of pig oocytes. Anim. Reprod. Sci. 111, 17–30.
| Involvement of calcium/calmodulin-dependent protein kinase kinase in meiotic maturation of pig oocytes.Crossref | GoogleScholarGoogle Scholar | 1:CAS:528:DC%2BD1cXhsFCgs7%2FL&md5=87dac586b8946d6b4a789e26367641dcCAS |
Yuan, S., Ortogero, N., Wu, Q., Zheng, H., and Yan, W. (2014). Murine follicular development requires oocyte DICER, but not DROSHA. Biol. Reprod. 91, 39.
| Murine follicular development requires oocyte DICER, but not DROSHA.Crossref | GoogleScholarGoogle Scholar |