Taxonomic tangles posed by human association – the urgent need for an evidence-based review of dingo and domestic dog taxonomy and nomenclature
Kylie M. Cairns A B * , Mike Letnic A B , Euan G. Ritchie C , Justin W. Adams D E F , Mathew S. Crowther
A B * , Mike Letnic A B , Euan G. Ritchie C , Justin W. Adams D E F , Mathew S. Crowther  G , Christopher R. Dickman
G , Christopher R. Dickman  G , Melanie Fillios H , Jack Pascoe I , Bradley P. Smith
G , Melanie Fillios H , Jack Pascoe I , Bradley P. Smith  J and Thomas M. Newsome G
J and Thomas M. Newsome G
A
B
C
D
E
F
G
H
I
J
Abstract
A serious challenge in modern taxonomy is whether, or how, relationships with humans have altered the evolutionary trajectory of species and how this should be dealt with in taxonomic nomenclature. Despite their historic treatment as a discrete taxon, the taxonomy and nomenclature of dingoes has been robustly debated for decades, in part due to their close evolutionary relationship to wolves and domestic dogs, as well as uncertainty about the role of humans in shaping their origins. One outcome of this ongoing scientific debate is a set of criteria regarding reproductive isolation, evolutionary distinctiveness and evolutionary origins that can be used to refine discussion about the appropriate taxonomy of dingoes. We review significant scientific advances in our knowledge of the distinct identity of dingoes in light of these criteria and discuss the need for scientists to assess the multi-stage process of domestication when considering species that may have interacted with humans over evolutionary timescales. Current evidence indicates that dingoes are on a different evolutionary trajectory to domestic dogs and thus we argue that the appropriate nomenclature for dingoes is Canis dingo or Canis lupus dingo, not Canis familiaris. We suggest that an independent evidence-based scientific review with expert consultation is one pathway through which taxonomic debate about dingoes and other controversial species could be resolved.
Keywords: Canis dingo, Canis familiaris, Canis lupus dingo, culturally significant species/entity, dingo, domestic dog, taxonomic nomenclature.
Introduction
Species concepts are a theoretical framework by which scientists attempt to classify and organise Earth’s biodiversity (Zachos 2016). A great challenge for modern taxonomy is how to deal with species or animal lineages that have complex evolutionary histories, or may have been associated with humans over time (Gentry et al. 2004; Clutton-Brock 2015). Here we use the term ‘association’ to describe a continuum of geographic co-occurrence and interaction between animal populations and humans that may or may not lead to domestication. This is because the intensity and occurrence of interactions with humans can, over long periods of time, lead to differences in morphology, behaviour or ecology between associated and non-associated or ‘wild’ animals. This is relevant to how we define a domestic lineage and whether domestic lineages, especially those that are genetically and morphologically distinct from their wild relatives, should be classed as separate species or excluded from taxonomic nomenclature (Clutton-Brock 1992; Gentry et al. 2004; Clutton-Brock 2015; Jackson et al. 2017; Zeller and Göttert 2019). It is therefore critical that scientists considering the taxonomic nomenclature of wild and domestic lineages use an accepted definition of domestication (Zeder 2012; Larson and Burger 2013; Lord et al. 2025).
It is currently common practice for lineages falling within the wild to domestic continuum to be distinguished using binomial taxonomic nomenclature (Gentry et al. 2004). This is reflected in International Commission on Zoological Nomenclature (ICZN) Opinion 2027 which allows ‘… taxonomic judgement as to what degree of domestication can be encompassed in the species-concept …’ (Gentry et al. 2004, p. 649). Distinguishing wild and domestic lineages is important to reflect differing evolutionary trajectories. Whilst some taxonomists hold the opinion that lineages associated with humans cannot speciate (Jackson et al. 2017; Zeller and Göttert 2019), this arbitrarily disregards the role natural selection plays in shaping lineages, especially those that may have had associations with humans but no deliberate artificial selection. It is also problematic to apply such ideas universally, given the complex relationships different peoples, including Indigenous peoples, have had and maintain with certain animals over millennia (Smith and Litchfield 2009; Fogg et al. 2015). For instance, Indigenous Australians connect to Country through a network of relationships which include kinship with Culturally Significant Species (Goolmeer et al. 2022). These linkages include very close relationships with Canis species which lived, and still live, entwined with those of Indigenous communities. Many indigenous languages have two words for Canis species that make a distinction between wild and human associated animals. Using nomenclature to differentiate between domestic and wild lineages also provides clarity to policy makers needing to conserve wildlife or identify illegal imports or exports of an endangered wild versus domestic species (Gentry et al. 2004; Clutton-Brock 2015; Zeller and Göttert 2019). In addition, what we call a species has conservation implications, particularly where classification embeds a taxon as a subspecies within common or widespread species with discordant management strategies, or where the taxonomic nomenclature may inappropriately preclude a lineage from legislative protection (Ritchie et al. 2018; vonHoldt et al. 2018).
Historically dingoes have been identified as a discrete taxon from domestic dogs, with three different species names in current widespread use (Crowther et al. 2014; Smith et al. 2019). In this paper, we focus on the evidence and knowledge gaps underlying debate on the taxonomy and nomenclature of dingoes and outline the consequences of taxonomic uncertainty for conservation policy. The three most common taxonomic names applied to dingoes over the last 30 years are: Canis lupus dingo (a subspecies of wolf), Canis dingo (a distinct species) and Canis familiaris (a domestic dog) (Daniels and Corbett 2003; Sillero-Zubiri 2009; Crowther et al. 2014; Allen et al. 2017; Jackson et al. 2017; Smith et al. 2019; Jackson et al. 2020; Cairns 2021; Shipman 2021; Krofel et al. 2022). The main points of difference in the use of these names revolve around the level of reproductive isolation between dingoes and domestic dogs, whether dingoes are distinct from wolves or extant domestic dogs and whether the ancestor of dingoes was domesticated, which some suggest precludes them from being considered a distinct taxon (Jackson et al. 2017).
With new scientific knowledge about the evolutionary and contemporary identity of dingoes, we suggest a pathway for resolving taxonomic debate about dingoes, which applies to other controversial species as well.
Background context: evolution of dogs
The evolution of dogs began approximately 20,000–40,000 years BP (Botigué et al. 2017; Freedman and Wayne 2017; Bergström et al. 2020). Current scientific knowledge suggests that rather than evolving directly from modern wolves, dogs evolved from either a common ancestor of the Holarctic wolf (Druzhkova et al. 2013; Thalmann et al. 2013; Freedman et al. 2014; Fan 2016; Freedman and Wayne 2017; Oetjens et al. 2018; Bergström et al. 2020; Perri et al. 2021; Gojobori et al. 2024) or originated directly from multiple geographically discrete grey wolf populations (Frantz et al. 2016; Freedman and Wayne 2017; Bergström et al. 2022). The exact nature and role of the relationship between early dogs and humans remains unclear. Within the Canis species complex, admixture between different Canis, and canid, lineages is common and may be adaptive (Sacks et al. 2013; vonHoldt et al. 2013; Skoglund et al. 2015; Fan et al. 2016; Frantz et al. 2016; Gopalakrishnan et al. 2018; Bergström et al. 2022; Gojobori et al. 2024). The diversification of early dogs into domestic dogs has been characterised by several periods of strong artificial selection, significantly predating and including during the transition of some human cultures to an agrarian lifestyle around 12,000 years ago (~10,000 BC) (Axelsson et al. 2013; Arendt et al. 2016; Cagan and Blass 2016; Freedman et al. 2016; Pendleton et al. 2018; Perri et al. 2021), and intensive selective breeding during modern breed formation around 200–500 years ago (vonHoldt et al. 2010; Parker et al. 2017).
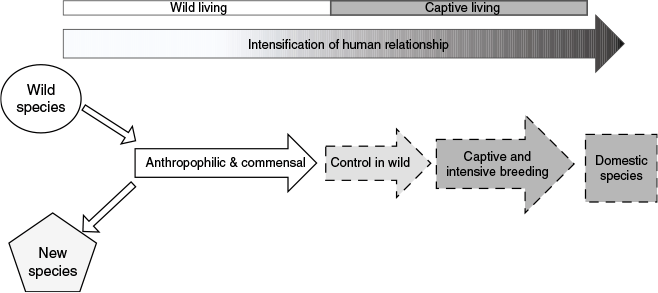
It is important not to conflate the original divergence of wolves and dogs within the genus Canis with domestication because: (1) these processes may not have occurred simultaneously; (2) may not have been originally human-directed; and (3) domestication itself is a multi-stage, fluid and long-term process (Zeder 2012; Larson and Burger 2013). Dogs are commonly posited to have become domesticated through a commensal pathway (Fig. 1) characterised by a slowly intensifying relationship with humans; progressing from anthropophilic to commensal to controlled breeding to companion animal (Driscoll et al. 2009; Zeder 2012; Larson and Burger 2013; Marshall et al. 2014; Zeller and Göttert 2019). Anthropophilic or commensal associations with humans, particularly if there is no or limited control of breeding, are rarely included in definitions of domestication (Lord et al. 2025). A key feature that most definitions of domestication rely upon is controlled and intensive breeding by humans following an intensification of the animal-human relationship. A new definition proposed by Lord et al. (2025) suggests domestic ‘should refer solely to those populations that are obligate synanthropes’ and are unable to form self-sustaining populations outside of anthropogenic niches.
Schematic of the ‘commensal’ domestication pathway originally hypothesised by Larson and Burger (2013) illustrating the intensification of the relationship with humans and changing relative strength of natural evolutionary forces. Solid lines indicate species or processes where natural evolutionary forces are occurring with limited impact of human selection on the lineage. Dashed lines indicate an increased impact and extent of human selection on the lineage to the exclusion of natural forces.
Resolving debate about the taxonomy of dingoes
Debate concerning the taxonomy and appropriate nomenclature for dingoes in Australia led Jackson et al. (2020, p. 352) to propose that one or more of the following conditions would need to be met for the dingo to be regarded a separate species:
New evidence of effective reproductive isolation of the dingo from related canids,
New genetic evidence that shows the dingo does not cluster within the diversity represented by domestic dog breeds,
New evidence that the dingo is an independently derived evolutionary lineage that is more distinct from domestic dog breeds, than these breeds are from each other,
In addition, the relationship of the Australian dingo population with other ‘dingo’ populations in mainland and island South-East Asia including New Guinea (Singing Dog) would need to be clarified.
Below, we demonstrate that there are multiple lines of evidence in support of these four criteria, and thus argue that dingoes should be considered a distinct taxonomic unit to domestic dogs, with discrete nomenclature. Whilst debate about taxonomic nomenclature is largely philosophical, it is important to note that nomenclature has a significant impact on the conservation and management of fauna and flora (Ritchie et al. 2018). Treatment of dingoes as a discrete species or sub-species enables governments to differentiate between dingoes and domestic dogs, allowing dingoes to be specifically protected or even listed as a threatened animal taxon. Nomenclature which does not distinguish between dingoes and domestic dogs constrains conservation and management efforts, effectively grouping dingoes with domestic dogs as the same entity.
Criterion 1: evidence of effective reproductive isolation
Molecular dating efforts indicate that dingoes diverged and have been genomically isolated from other Canis lineages, including wolves and domestic dogs, for 8000–11,000 years (Table 1; Cairns and Wilton 2016; Bergström et al. 2020; Zhang et al. 2020; Bergström et al. 2022; Souilmi et al. 2024; Weeks et al. 2024). Carbon dating of archaeological dingo remains from southwestern Australia indicates that they have been widespread across Australia for at least 3500 years (Table 1; Balme et al. 2018).
| Category | Source | Data type | Method/approach | Estimated divergence time (years BP) | Estimated arrival in Australia (years BP) | Key findings | |
|---|---|---|---|---|---|---|---|
| Archaeological evidence | Balme et al. (2018) | Fossil (dingo bone remains) | Direct Accelerator Mass Spectrometry (AMS) radiocarbon dating from Madura Cave | – | 3348–3081 | Provides the oldest directly dated dingo remains; firm lower boundary for dingo arrival | |
| Morphological evidence | Koungoulos et al. (2024) | Morphology (crania + postcrania) | 3D geometric morphometrics + direct AMS dating | – | 3000–3300 | Shows early southeastern dingoes had smaller builds and closer cranial affinity to East Asian and New Guinea Singing Dogs (NGSD); evidence of early regional variation | |
| Genetic evidence – mitochondrial DNA | Savolainen et al. (2004) | mtDNA (control region) | Sequencing + phylogenetic distance to founder haplotype A29 | ~5000 (range 4600–10,800) | ~5000 (same range used as proxy) | All dingo mtDNA types cluster around A29; suggests a single introduction event and small founder population from East Asia | |
| Oskarsson et al. (2012) | mtDNA (control region) | Broader analysis of SE Asian, Polynesian, and Australian dogs | ~4600–18,300 | ~4000–8000 (implied by mtDNA spread and diversity) | Identifies route through Mainland Southeast Asia; dingo haplotype diversity suggests older introduction than archaeological record alone. Dingo haplotypes are absent from Taiwan and the Philippines, suggesting a mainland Southeast Asia route, not Austronesian expansion | ||
| Cairns and Wilton (2016) | Whole mtDNA +13 nuclear loci | Bayesian divergence time estimation | ~8300 | 6000–10,000 | Finds evidence of two distinct dingo populations; plausible multiple introductions and regional structuring | ||
| Genetic evidence – genome-wide | Zhang et al. (2020) | Whole genome + mtDNA | Phylogenetics + demographic modelling | ~8300 | – | Indicates divergence from southern East Asian dogs; identifies selection in dietary and behavioural genes related to wild living | |
| Kumar et al. (2023) | Whole genome | Genomic diversity, runs of homozygosity (ROH), deleterious load | ~4000–11,000 | – | Finds strong founder effects and lowest diversity among canids; supports ancient colonisation and long isolation | ||
| Genetic evidence – ancient DNA | Souilmi et al. (2024) | Ancient and modern genome-wide | mtDNA, nuclear DNA, demographic modelling | mtDNA ancestor ~3000–8000 | Ancient samples: 400–2746 | Demonstrates deep population structure already existed thousands of years ago; confirms continuity and minimal post-colonial admixture | |
| Synthesis and interpretive reviews | Fillios and Taçon (2016) | Genetic + archaeological evidence (from prior studies) | Critical synthesis of archaeological and genetic data | – | 3500–10,000 | Integrates multiple data types; proposes Sulawesi as parsimonious origin; supports early, possibly multiple arrivals | |
| Smith et al. (2019) | Integrative review (fossil, genetic, morphological, behavioural) | Critical synthesis of archaeological, molecular, phenotypic, ecological and taxonomic data | ~5000–10,000 | 3500–10,000 | Reviews multiple lines of evidence supporting dingo divergence from East Asian dogs prior to agriculture. Affirms arrival in Australia at least 3500 years ago, likely between 5000–10,000 BP. Suggests arrival was independent of Neolithic cultural markers, possibly with hunter-gatherers. Highlights long isolation from other canids post-arrival | ||
| Cairns (2021) | Genome-wide single nucleotide polymorphism (SNP) data (from prior studies); functional gene analysis (e.g. AMY2B) | Review and reinterpretation of existing genomic data; emphasis on functional traits and divergence context | ~8000–10,000 | ≥3500 | Argues dingoes diverged from domestic dogs before the rise of agriculture; highlights AMY2B deletion (lack of starch adaptation); supports early, non-Neolithic separation and long isolation from other dogs |
Note: some genome-wide studies estimate dingo divergence from other dog lineages at 8000–11,000 BP (e.g. Zhang et al. 2020; Kumar et al. 2023), while archaeological evidence supports arrival in Australia at least ~3500–4000 BP (Balme et al. 2018). It is possible that genetic divergence occurred prior to arrival and in Southeast Asia or near Oceania, not within Australia.
High levels of genetic sharing between dingoes and Japanese wolves (Gojobori et al. 2024) suggest that the dingo’s ancestor may have been an Asiatic wolf, however ancient DNA studies with broader sampling of Asiatic wolves are needed to confirm this. A strict biological species concept interpretation of canid taxonomy is problematic due to the widespread occurrence of admixture, which may be evolutionarily adaptive, between Canis lineages (Gopalakrishnan et al. 2018; Schweizer 2018; Smith et al. 2019; vonHoldt and Aardema 2020; Pilot 2021). However, whilst many Canis species, including jackals (Canis spp.), coyotes (Canis latrans) and wolves (Canis spp.) remain biologically compatible, behavioural and geographic barriers to reproduction act to minimise geneflow between species (or subspecies) and maintain effective reproductive isolation. In this context complete reproductive isolation is not expected to be present between Canis species that carry the same chromosome complement.
There have been significant changes to our knowledge about the genetic identity of dingoes and patterns of interspecific hybridisation with domestic dogs. Cairns et al. (2023) demonstrated that there is limited contemporary interbreeding between dingoes and domestic dogs across the Australian landscape, revealing effective reproductive isolation between the lineages. Weeks et al. (2024) corroborated the finding of limited contemporary gene flow between dingoes and modern domestic dogs, especially in Victoria. Additionally, Kreplins et al. (2024) showed little dingo introgression in Aboriginal community dogs in remote Western Australia, even though the dingoes and domestic dogs lived in close proximity. Ancient DNA studies have confirmed the divergent pre-historic and present-day identity of dingoes in Oceania from modern domestic dogs (Souilmi et al. 2024; Scarsbrook et al. In Press). While Scarsbrook et al. (In Press) report historical geneflow (1800s–1960s) from domestic dogs into southeastern dingoes, they find evidence that dog alleles retained in contemporary dingoes have been positively selected and may have played a role in mitigating inbreeding depression. Together these studies highlight that contemporary dingo populations have maintained their evolutionary distinctiveness from domestic dogs, with limited present-day geneflow between the lineages.
There are several mechanisms possibly driving reproductive isolation between dingoes and modern domestic dog lineages in Australia. There is limited opportunity for dingoes and domestic dogs to interact with over 85% of the human (and domestic dog) population residing in urban areas/capital cities within 50 km of the coast (Australian Bureau of Statistics 2020). Further, around 80% of pet domestic dogs in Australia are neutered (Animal Medicines Australia 2019), limiting opportunity for reproduction between dingoes and domestic dogs. There are also key differences in the reproductive strategies of dingoes (seasonal breeding and biparental care) and domestic dog lineages (non-seasonal breeding and uniparental care) (Lord et al. 2013; Cursino et al. 2017). As such, female dingoes are only able to conceive during a short window, limiting the opportunity for hybridisation. Furthermore, strong social behaviours, such as the maintenance of cohesive family pack structures and territories (Thomson et al.1992), reduces the likelihood of mating between an established dingo family and interloping dogs.
One key point of difference between dingoes and domestic dogs is that dingoes can live independently in the wild. The absence of wild domestic dog populations in Australia (Stephens et al. 2015; Cairns et al. 2021; Stephens et al. 2022) suggests that modern domestic dogs are not capable of living independently, forming self-sustaining populations in the wild or establishing themselves within the pack structure of dingoes; which may be driven by an inability to meet their own energetic needs or reproduce successfully. This inability of domestic dogs to establish populations in the wild is made particularly apparent by the absence of wild dog populations in Tasmania and New Zealand (Letnic et al. 2012; King and Forsyth 2021). Both of these islands support a similar range of dog breeds and land-management strategies to mainland Australia, but both lack wild dogs. The best explanation for the presence of dingoes on mainland Australia and the absence of wild domestic dogs across mainland Australia, Tasmania and New Zealand is that dingoes and their hybrids have the genetic material necessary to live in the wild but domestic dogs do not. The inability to self-sustain in Australian ecosystems reproductively isolates domestic dogs from dingoes and highlights that while domestic dogs are obligate synanthropes, dingoes are not (Lord et al. 2025). Reproductive isolation between dingoes and domestic dogs may be partly due to pre-zygotic behavioural barriers as well as posited differences in ecological and reproductive fitness (Letnic et al. 2012; Ballard and Wilson 2019; King and Forsyth 2021).
Criterion 2: evidence that dingoes are genetically distinct from domestic dog breeds
Genomic studies find evidence of at least three lineages of dog that diverged from ancestral wolf populations: eastern dogs, arctic dogs and western dogs (Frantz et al. 2016; Bergström et al. 2020, 2022; Feuerborn et al. 2021; Perri et al. 2021; Gojobori et al. 2024). These three early dog lineages either evolved from different wolf populations (Frantz et al. 2016; Freedman and Wayne 2017; Bergström et al. 2022) or diverged from a common protodog ancestor prior to the rise of agriculture and the initiation of intensive domestication processes (Arendt et al. 2016; Bergström et al. 2020; Zhang et al. 2020; Cairns 2021; Perri et al. 2021; Field et al. 2022; Gojobori et al. 2024).
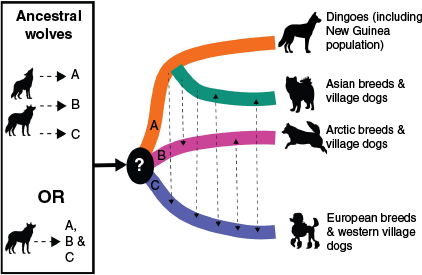
Dingoes are a wild-living canid descended from the eastern dog lineage whilst extant modern domestic dogs primarily originate from the western and Arctic dog lineages (Fig. 2; Bergström et al. 2020, 2022; Gojobori et al. 2024). While village dogs found in Asia today carry ancestry from the eastern dog lineage, they have experienced ongoing geneflow from western dog lineages over the last few thousand years (Fig. 2; Pilot et al. 2015; Shannon et al. 2015; Bergström et al. 2020; Surbakti et al. 2020; Ahn et al. 2023; Gojobori et al. 2024). Both ancient and modern western dogs are more closely related to the Arctic dog lineage than to eastern dogs, including the dingo (Bergström et al. 2020; Surbakti et al. 2020; Feuerborn et al. 2021; Perri et al. 2021; Bergström et al. 2022). Modern domestic dog breeds such as the Siberian husky share ancestry with the ancient Arctic dog lineage, but they also carry ancestry from western dogs and have been selectively bred by humans over the last few thousand years (Brown et al. 2013; Sinding et al. 2020; Feuerborn et al. 2021; Perri et al. 2021). Once the ancestor of dingoes arrived in Oceania the lineage has remained isolated from other Canis species including domestic dogs (Ardalan et al. 2012; Oskarsson et al. 2012; Sacks et al. 2013; Cairns and Wilton 2016; Cairns et al. 2018; Greig et al. 2018; Zhang et al. 2020; Cairns 2021).
Schematic of the hypothesised evolutionary origins of dogs and dingoes. Wolves are the ancestors of the first dogs, however, it is unclear whether a single or multiple wolf populations were the ancestors of early dogs. Following the divergence of dogs from wolves, there were three dog lineages, (a) the Eastern or Asian dog lineage, (b) the Arctic dog lineage and (c) the West Eurasian dog lineage. The evolutionary distinctiveness of dingoes is highlighted by an extended period of isolation in Oceania, while geneflow occurred extensively between other dog lineages across the globe during the last 10,000 years, as depicted by dashed black lines. Figure drawn in Adobe Illustrator 2025 based on our current knowledge of the evolutionary origins of dingoes and dogs including the work of Pilot et al. (2015), Shannon et al. (2015), Parker et al. (2017), Bergström et al. (2020), Surbakti et al. (2020), Zhang et al. (2020), Cairns (2021), Bergström et al. (2022), Cairns et al. (2023), Gojobori et al. (2024).
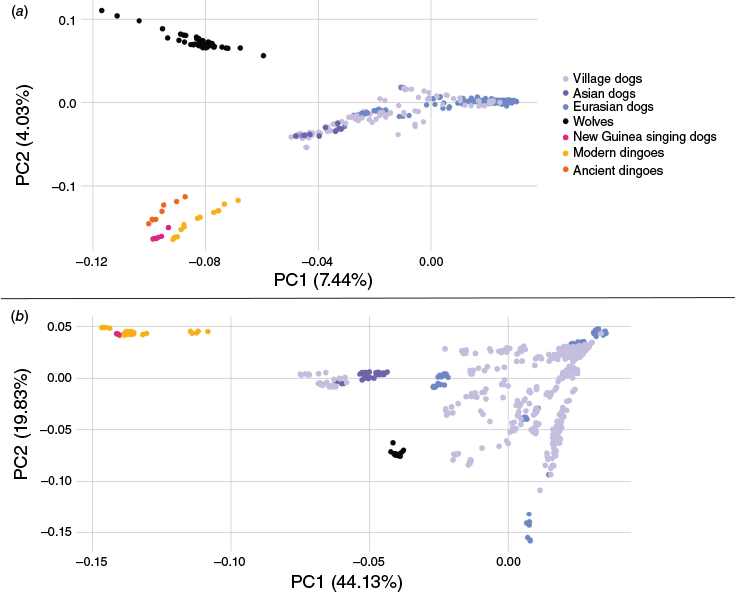
Ancient DNA studies have corroborated the distinctiveness of modern and ancient dingoes from extant domestic dogs (Fig. 3; Souilmi et al. 2024). Genome-wide marker studies have demonstrated strong evidence of genetic differentiation between dingoes, domestic dogs and wolves (Fig. 3; Cairns et al. 2023; Weeks et al. 2024). Cairns et al. (2023) reported high levels of genetic differentiation between dingoes and domestic dogs, with calculated Fixation Index (FST) values exceeding 0.25. Genomic comparisons between dingoes, domestic dogs and wolves have identified evidence of evolutionary change since their arrival in Australia (Zhang et al. 2020; Field et al. 2022). Field et al. (2022) reported the presence of structural chromosomal differences (inversions and duplications) between wolves, dingoes and domestic dogs. Furthermore, genomic studies found that dingoes were not subjected to the same intensive selective breeding and dietary modifications as the ancestors of modern domestic dog breeds (Arendt et al. 2016; Zhang et al. 2020; Field et al. 2022).
(a) Principal Component Analysis (PCA) based on nuclear genomic data of dingoes (modern and ancient), New Guinea singing dogs, wolves and global domestic dog populations. The PCA has been re-plotted in R using ggplot2 and SMARTPCA analysis from Souilmi et al. (2024). Note that we have excluded two ancient dingo samples from Souilmi et al. (2024), one that was sequenced using a domestic dog bait capture method and one which had extremely low coverage (<0.01×). (b) PCA from Cairns (2021) based upon filtered Illumina genome wide Single Nucleotide Polymorphism (SNP) data (113,523 sites) from 657 canid samples including wolves, dingoes, New Guinea singing dogs and global dog populations, replotted in R using ggplot2.
Criterion 3: evidence that dingoes form an independently derived evolutionary lineage that is more distinct from domestic dog breeds than dog breeds are from each other
The direct ancestors and processes underlying the origins of early dogs remain unclear, including whether there was a single or multiple ancestral wolf populations (Freedman and Wayne 2017; Bergström et al. 2020; Perri et al. 2021; Bergström et al. 2022; Gojobori et al. 2024). However, there is a wide range of morphological and physiological evidence that dingoes are distinct from domestic dog lineages and are more divergent from domestic dog lineages than breeds are from each other (Crowther et al. 2014; Smith et al. 2019; Cairns 2021; Smith et al. 2023). Modern genomic evidence confirms the distinctiveness of modern dingoes from extant domestic dogs (Bergström et al. 2020; Surbakti et al. 2020; Zhang et al. 2020; Cairns 2021; Field et al. 2022; Cairns et al. 2023; Gojobori et al. 2024). Both modern and ancient dingoes (including the New Guinea Singing Dog (NGSD)) form a distinct genetic cluster to wolves as well as domestic dogs (Fig. 3).
Dingoes, including the NGSD, are an example of the eastern dog lineage which diverged early and has remained free from admixture with other wild canids or dog lineages (Bergström et al. 2020; Surbakti et al. 2020; Gojobori et al. 2024). The ancestry of extant domestic dogs is complex due to historical admixture and migration across the globe as well as the actions of humans in selective breeding (Gopalakrishnan et al. 2018; Bergström et al. 2022). Extant domestic dog breeds and village dog populations in Eurasia, Europe, Africa and the Americas descend almost exclusively from the western dog lineage or from admixture between various dog lineages (Pilot et al. 2015; Shannon et al. 2015; Bergström et al. 2020; Surbakti et al. 2020). Furthermore, most extant domestic dog breeds were only developed in the last 200–400 years and are genetically shallow lineages (Parker et al. 2017). Based on this genomic evidence, dingoes represent an evolutionary lineage that is independent and discrete from extant domestic dogs and has been so for thousands of years.
Some authors argue that because the ancestor of dingoes are likely to have been transported to Oceania by humans, then they are a domesticate and are not evolutionarily independent from domestic dogs (Jackson et al. 2017; Jackson et al. 2019; Jackson et al. 2020). However, humans have transported many animal lineages around the globe and these lineages are not considered domesticates, according to conventional definitions of domestication (Heinsohn 2014; Ahlgren et al. 2016; Hofman and Rick 2018; Lord et al. 2025). Good examples of this are rusa deer (Cervus timorensis) and Malay civets (Viverra tangalunga), which are believed to have been transported by humans across Wallace’s line during the last 5000–10,000 years (Veron et al. 2014; Martins et al. 2018). Like dingoes in Australia, rusa deer and Malay civits live in the wild on numerous islands and are not considered to be a domesticate.
Further evidence that dingoes are a distinct lineage from domestic dogs is their lack of AMY2B copy number variation, which is a dietary adaptation observed in ancient and modern domestic dogs associated with prehistoric agricultural expansion (Arendt et al. 2016). The dingo lineage diverged from its ancestors roughly 8000–11,000 years ago (Cairns and Wilton 2016; Bergström et al. 2020; Zhang et al. 2020; Souilmi et al. 2024), which was likely prior to the intensification of the human-canine relationship, particularly human control of canine reproduction and selective breeding. Studies of Y-chromosome and mitochondrial DNA indicate that dingoes reflect an different older migration event than the ancestors of dogs in Island South East Asia and the Pacific (Sacks et al. 2013; Greig et al. 2018). Thus, in our opinion it is erroneous to label all dog lineages as domesticates when not all dog lineages had, or were posited to have, a relationship with humans that falls within domestication criteria (Fig. 1; Manwell and Baker 1984; Galibert et al. 2011; Zeder 2012; Larson and Burger 2013; Smith et al. 2019; Cairns 2021; Purugganan 2022; Smith et al. 2023). Whilst early dogs and the ancestor of dingoes may have been commensal or anthropophilic, in Oceania dingoes live (or lived) free from human interaction or control. We know that some dingoes lived with or in close association to Indigenous Australians (Smith and Litchfield 2009; Koungoulos and Fillios 2020; Koungoulos 2021), but it is unclear what evolutionary impact this had on free-living dingoes as a population.
Criterion 4: relationship of dingoes to dogs from south east Asia
It has been commonly postulated that village dogs in south east Asia with phenotypic similarity to dingoes are either the direct progenitors of dingoes, or are dingoes themselves (Corbett 2001). However, modern genomic evidence has illustrated that dingoes are distinct from extant Asian village dogs (Fig. 3; Surbakti et al. 2020; Field et al. 2022; Cairns et al. 2023; Kumar et al. 2023; Gojobori et al. 2024; Souilmi et al. 2024). Indeed, Ahn et al. (2023) found that dingoes (and NGSD) form a distinct population to Asian village dogs and exhibit FST values of >0.25, similar to the FST values observed between wolves and Asian village dogs.
Extant Asian village dogs are a heavily admixed lineage, carrying small amounts of persisting eastern dog lineage, as well as significant ancestry from western and modern domestic dog lineages from recurring admixture over the last several thousand years (Fig. 2; Pilot et al. 2015; Shannon et al. 2015; Bergström et al. 2020; Surbakti et al. 2020; Ahn et al. 2023; Gojobori et al. 2024). Scientists should be cautious in interpreting phylogenetic analyses depicting Asian village dogs or breeds as basal to dingoes because many phylogenetic methods inadequately account for patterns of ancient, historical and modern admixture (Surbakti et al. 2020; Bergström et al. 2022). It is indisputable that dingoes and early Asian dogs share ancestors; however, the evolutionary trajectory of dingoes has diverged due to adaptation to Australian environments and isolation from other wild or domestic canids. Asian dogs, on the other hand, have continued along a domestication trajectory, living in close proximity to humans (obligate synanthropes) and experiencing persistent genetic swamping from Western dogs.
Conclusions
Significant advances in genomic research provide robust evidence that dingoes are on a biologically meaningful and divergent evolutionary trajectory from both domestic dogs and extant populations of wolves. Our existing knowledge of morphological and biological differentiation between dingoes, wolves and domestic dogs (Crowther et al. 2014; Smith et al. 2019), along with the new evidence of reproductive isolation (Cairns et al. 2023; Weeks et al. 2024), highlights the need for a reconsideration of the taxonomic status and nomenclature used for dingoes. We disagree with the interpretation of some authors that any association with humans or domestication pressure (passive or active), at any time point, precludes lineages from nomenclatural distinction (Jackson et al. 2017; Gaunitz et al. 2018; Jackson et al. 2019; Jackson et al. 2020). This interpretation is problematic because it precludes any anthropophilic or commensal lineage from being defined as a species, no matter how distinct the lineage has become or how long the lineage has been distinct.
When considering how we make these distinctions it is also imperative to consider the cultural context, acknowledging traditional knowledge about the identity of species and relationships between a species and humans. In the case of dingoes, human-dingo relationships are sacred, and dingoes prominently feature in the Law/Lore of Indigenous Australians, as well as being a part of the network of relationships in Country. We argue that for ongoing discussions about the nomenclature or taxonomy of species where humans may have played a role, it is important that researchers properly define the human-animal relationship (i.e. domestication), as well as highlighting the relative impact and extent of human selection on the lineage to the exclusion of natural forces (Fig. 1).
We concur with Gentry et al. (2004) that nomenclatural differentiation between domestic and wild lineages should be adopted to provide clarity to legislators, wildlife managers and applied scientists (conservation, ecology, behaviour etc.). There are several ways that taxonomic nomenclature rules and conventions can be applied to dingoes. It is clear from their historical treatment and emerging genomic evidence that dingoes represent a distinct taxonomic unit to domestic dogs, what is less clear is whether dogs, including both dingoes and domestic dogs, are a discrete species from wolves. If it is agreed that dogs are a discrete species from wolves and accepting ICZN Opinion 2027 that wild and domestic lineages should have distinct nomenclature, we could utilise Canis dingo for the dingo lineage and Canis familiaris to refer to domestic lineage(s) of dogs. An alternative approach utilising trinomial nomenclature would be to nominate nomenclature for the original protodog(s) and then designate dingoes and domestic dogs as subspecies of that dog lineage. Either of these solutions allows taxonomic end-users to unambiguously distinguish between distinct evolutionary lineages of dog – one which is a valid wild taxon and the other which is a domestic complex. These approaches acknowledge the shared evolutionary history of these lineages, their differentiation from wolves and the divergent evolutionary trajectories of dingoes and domestic dogs. If the taxonomic differentiation of dogs from wolves is considered unclear then an alternative solution would be to continue including dingoes and dogs under Canis lupus, referring to the distinct evolutionary lineages as Canis lupus dingo and Canis lupus familiaris. We contend that a binomial or trinomial variant of Canis familiaris should not be applied to dingoes given this nomenclature refers specifically to modern domestic dogs.
Conservation implications of taxonomic uncertainty for dingoes
The taxonomic treatment and nomenclature of dingoes is not simply philosophical but has ‘real world’ practical implications in terms of conservation and management. While the Australian Mammal Taxonomy Consortium argues that ‘naming the dingo as a separate species should not be necessary to protect it …’, the reality is that if a lineage is not considered taxonomically discrete at either the species or subspecies level then legislative protection is difficult. Indeed, the New South Wales (NSW) Threatened Species Scientific Committee decided in May 2024 that due to the Australian Faunal Directory listing dingoes as Canis familiaris, ‘the dingo is not eligible for listing under the Act as it is not recognised as a defined subspecies’ (NSW Threatened Species Scientific Committee, pers. comm.). This difficulty is based on the assessment that if dingoes share the nomenclature Canis familiaris with domestic dogs, then they are not evolutionarily or taxonomically distinct, and as such would not warrant specific conservation management. This also has implications for the status of dingoes, and plausibly domestic dogs, as a native animal under Australian legislation. Currently, the federal government and most states classify dingoes legislatively as a native animal based on their presence in Australia prior to 1400 AD (or prior to 1788), but if dingoes and domestic dogs are not taxonomically discrete then this may also unintentionally include domestic dogs.
Conversely, taxonomic treatment of dingoes as Canis lupus dingo or Canis dingo allows jurisdictions to distinguish between evolutionarily distinct dingoes and domestic dogs under legislation (and in public policy), thus practically enabling different conservation and management policies for dingoes and domestic dogs. This differentiation also would not inappropriately include dingoes in companion animal legislation. Furthermore, taxonomic treatment of dingoes as either a species or subspecies would allow appropriate acknowledgement and management of the dingo as a ‘culturally significant species/entity’ (Goolmeer et al. 2022) without inappropriately including domestic dogs.
Next steps
It is unlikely that debate about the taxonomy and nomenclature of dingoes will be resolved by ongoing academic debate in the literature. The ICZN (1999) specifically notes in the Code’s introduction that (1) ‘the Code refrains from infringing upon taxonomic judgment …’ and (2) ‘nomenclature does not determine the inclusiveness or exclusiveness of any taxon … but rather provides the name that is to be used for a taxon whatever taxonomic limits and rank are given to it’. There is academic freedom in how scientists define species, how species are distinguished from subspecies and how nomenclature is applied to taxonomy (Gentry et al. 2004; De Queiroz 2007; Clutton-Brock 2015; Thiele et al. 2021). However, for end-users, and those outside of, or not versed in the nuances of academic debate, ongoing taxonomic debate results in instability (Coates et al. 2018). Any proposal to seek consensus on the naming of dingoes needs to engage in broad consultation with taxonomists, taxon specific experts as well as seeking feedback from both Australian and international scientific organisations, and representatives of First Nations, to inform nomenclature decisions; i.e. the International Union for the Conservation of Nature (IUCN) Canid Specialist Group and the Australian Mammal Society. Scientists communicating with policy makers and wildlife managers should ensure that decisions about taxonomic nomenclature are transparent, evidence-based and presented as scientific hypotheses with alternative views acknowledged. We further propose an approach similar to that employed by the United States National Academies of Sciences, Engineering and Medicine where a comprehensive, independent and evidence-based review of the taxonomic status of red wolves (Canis rufus) and Mexican wolves (Canis lupus baileyi) was undertaken including consultation with taxonomic experts and wolf experts with diverse opinions (National Academies of Sciences, Engineering, and Medicine 2019). Given significant scientific progress in recent years, and the implications for public policy, we aim to advance this discussion and advocate for an independent review of the taxonomic status and nomenclature of dingoes by an appropriate Australian institution such as the Australian Academy of Science.
Conflicts of interest
K. M. C. is co-coordinator of the IUCN Species Survival Commission (SSC) Canid Specialist Group’s Dingo Working Group and a voluntary scientific advisor to the Australian Dingo Foundation and New Guinea Highland Wild Dog Foundation. B. P. S. is a member of the IUCN SSC Canid Specialist Group’s Dingo Working Group and a non-financial director of the Australian Dingo Foundation. The other authors declare no conflicts of interest.
Acknowledgements
Dingoes are considered kin by many First Nations peoples across mainland Australia, and have deep connections with Indigenous culture and Country. We would like to acknowledge and pay our respects to these connections.
References
Ahlgren, H., Norén, K., Angerbjörn, A., and Lidén, K. (2016). Multiple prehistoric introductions of the mountain hare (Lepus timidus) on a remote island, as revealed by ancient DNA. Journal of Biogeography 43(9), 1786-1796.
| Crossref | Google Scholar |
Ahn, B., Kang, M., Jeon, H., Kim, J.-S., Jiang, H., Ha, J., and Park, C. (2023). Origin and population structure of native dog breeds in the Korean peninsula and East Asia. iScience 26, 106982.
| Crossref | Google Scholar | PubMed |
Allen, B. L., Allen, L. R., Ballard, G., Jackson, S. M., and Fleming, P. J. S. (2017). A roadmap to meaningful dingo conservation. Canid Biology and Conservation 20(11), 45-56.
| Google Scholar |
Animal Medicines Australia (2019). Pets in Australia: A national survey of pets and people, Animal Medicines Australia. Available at https://animalmedicinesaustralia.org.au/resources/pets-in-australia-a-national-survey-of-pets-and-people
Ardalan, A., Oskarsson, M., Natanaelsson, C., Wilton, A. N., Ahmadian, A., and Savolainen, P. (2012). Narrow genetic basis for the Australian dingo confirmed through analysis of paternal ancestry. Genetica 140, 65-73.
| Crossref | Google Scholar | PubMed |
Arendt, M., Cairns, K. M., Ballard, J. W. O., Savolainen, P., and Axelsson, E. (2016). Diet adaptation in dog reflects spread of prehistoric agriculture. Heredity 117, 301-306.
| Crossref | Google Scholar | PubMed |
Axelsson, E., Ratnakumar, A., Arendt, M.-L., Maqbool, K., Webster, M. T., Perloski, M., Liberg, O., Arnemo, J. M., Hedhammar, A., and Lindblad-Toh, K. (2013). The genomic signature of dog domestication reveals adaptation to a starch-rich diet. Nature 495(7441), 360-364.
| Crossref | Google Scholar | PubMed |
Ballard, J. W. O., and Wilson, L. A. B. (2019). The Australian dingo: untamed or feral? Frontiers in Zoology 16, 2.
| Crossref | Google Scholar | PubMed |
Balme, J., O’Connor, S., and Fallon, S. (2018). New dates on dingo bones from Madura Cave provide oldest firm evidence for arrival of the species in Australia. Scientific Reports 8(1), 9933.
| Crossref | Google Scholar | PubMed |
Bergström, A., Frantz, L., Schmidt, R., Ersmark, E., Lebrasseur, O., Girdland-Flink, L., Lin, A. T., Storå, J., Sjögren, K.-G., Anthony, D., Antipina, E., Amiri, S., Bar-Oz, G., Bazaliiskii, V. I., Bulatović, J., Brown, D., Carmagnini, A., Davy, T., Fedorov, S., Fiore, I., Fulton, D., Germonpré, M., Haile, J., Irving-Pease, E. K., Jamieson, A., Janssens, L., Kirillova, I., Horwitz, L. K., Kuzmanovic-Cvetković, J., Kuzmin, Y., Losey, R. J., Dizdar, D. L., Mashkour, M., Novak, M., Onar, V., Orton, D., Pasarić, M., Radivojević, M., Rajković, D., Roberts, B., Ryan, H., Sablin, M., Shidlovskiy, F., Stojanović, I., Tagliacozzo, A., Trantalidou, K., Ullén, I., Villaluenga, A., Wapnish, P., Dobney, K., Götherström, A., Linderholm, A., Dalén, L., Pinhasi, R., Larson, G., and Skoglund, P. (2020). Origins and genetic legacy of prehistoric dogs. Science 370(6516), 557.
| Crossref | Google Scholar | PubMed |
Bergström, A., Stanton, D. W. G., Taron, U. H., Frantz, L., Sinding, M. S., Ersmark, E., Pfrengle, S., Cassatt-Johnstone, M., Lebrasseur, O., Girdland-Flink, L., Fernandes, D. M., Ollivier, M., Speidel, L., Gopalakrishnan, S., Westbury, M. V., Ramos-Madrigal, J., Feuerborn, T. R., Reiter, E., Gretzinger, J., Münzel, S. C., Swali, P., Conard, N. J., Carøe, C., Haile, J., Linderholm, A., Androsov, S., Barnes, I., Baumann, C., Benecke, N., Bocherens, H., Brace, S., Carden, R. F., Drucker, D. G., Fedorov, S., Gasparik, M., Germonpré, M., Grigoriev, S., Groves, P., Hertwig, S. T., Ivanova, V. V., Janssens, L., Jennings, R. P., Kasparov, A. K., Kirillova, I. V., Kurmaniyazov, I., Kuzmin, Y. V., Kosintsev, P. A., Lázničková-Galetová, M., Leduc, C., Nikolskiy, P., Nussbaumer, M., O’Drisceoil, C., Orlando, L., Outram, A., Pavlova, E. Y., Perri, A. R., Pilot, M., Pitulko, V. V., Plotnikov, V. V., Protopopov, A. V., Rehazek, A., Sablin, M., Seguin-Orlando, A., Storå, J., Verjux, C., Zaibert, V. F., Zazula, G., Crombé, P., Hansen, A. J., Willerslev, E., Leonard, J. A., Götherström, A., Pinhasi, R., Schuenemann, V. J., Hofreiter, M., Gilbert, M. T. P., Shapiro, B., Larson, G., Krause, J., Dalén, L., and Skoglund, P. (2022). Grey wolf genomic history reveals a dual ancestry of dogs. Nature 607(7918), 313-320.
| Crossref | Google Scholar | PubMed |
Botigué, L. R., Song, S., Scheu, A., Gopalan, S., Pendleton, A. L., Oetjens, M., Taravella, A. M., Seregély, T., Zeeb-Lanz, A., Arbogast, R.-M., Bobo, D., Daly, K., Unterländer, M., Burger, J., Kidd, J. M., and Veeramah, K. R. (2017). Ancient European dog genomes reveal continuity since the Early Neolithic. Nature Communications 8(1), 16082.
| Crossref | Google Scholar | PubMed |
Brown, S. K., Darwent, C. M., and Sacks, B. N. (2013). Ancient DNA evidence for genetic continuity in Arctic dogs. Journal of Archaeological Science 40(2), 1279-1288.
| Crossref | Google Scholar |
Cagan, A., and Blass, T. (2016). Identification of genomic variants putatively targeted by selection during dog domestication. BMC Ecology and Evolution 16, 10.
| Crossref | Google Scholar | PubMed |
Cairns, K. M. (2021). What is a dingo – origins, hybridisation and identity. Australian Zoologist 41(3), 322-337.
| Crossref | Google Scholar |
Cairns, K. M., and Wilton, A. N. (2016). New insights on the history of canids in Oceania based on mitochondrial and nuclear data. Genetica 144(5), 553-565.
| Crossref | Google Scholar | PubMed |
Cairns, K. M., Shannon, L. M., Koler-Matznick, J., Ballard, J. W. O., and Boyko, A. R. (2018). Elucidating biogeographical patterns in Australian native canids using genome wide SNPs. PLoS One 13(6), e0198754.
| Crossref | Google Scholar | PubMed |
Cairns, K. M., Crowther, M. S., Nesbitt, B., and Letnic, M. (2021). The myth of wild dogs in Australia: are there any out there? Australian Mammalogy 44, 67-75.
| Crossref | Google Scholar |
Cairns, K. M., Crowther, M. S., Parker, H. G., Ostrander, E. A., and Letnic, M. (2023). Genome-wide variant analyses reveal new patterns of admixture and population structure in Australian dingoes. Molecular Ecology 32, 4133-4150.
| Crossref | Google Scholar | PubMed |
Clutton-Brock, J. (1992). The process of domestication. Mammal Review 22(2), 79-85.
| Crossref | Google Scholar |
Coates, D. J., Byrne, M., and Moritz, C. (2018). Genetic Diversity and Conservation Units: Dealing With the Species-Population Continuum in the Age of Genomics. Frontiers in Ecology and Evolution 6, 165.
| Crossref | Google Scholar |
Crowther, M. S., Fillios, M., Colman, N., and Letnic, M. (2014). An updated description of the Australian dingo (Canis dingo Meyer, 1793). Journal of Zoology 293, 192-203.
| Crossref | Google Scholar |
Cursino, M. S., Harriott, L., Allen, B. L., Gentle, M., and Leung, L. K. P. (2017). Do female dingo–dog hybrids breed like dingoes or dogs? Australian Journal of Zoology 65(2), 112-119.
| Google Scholar |
Daniels, M. J., and Corbett, L. (2003). Redefining introgressed protected mammals: when is a wildcat a wild cat and a dingo a wild dog? Wildlife Research 30(3), 213-218.
| Crossref | Google Scholar |
De Queiroz, K. (2007). Species Concepts and Species Delimitation. Systematic Biology 56(6), 879-886.
| Crossref | Google Scholar | PubMed |
Driscoll, C. A., Macdonald, D. W., and O’brien, S. J. (2009). From wild animals to domestic pets, an evolutionary view of domestication. Proceedings of the National Academy of Sciences of the United States of America 106 Suppl 1(Supplement 1), 9971-8.
| Crossref | Google Scholar | PubMed |
Druzhkova, A. S., Thalmann, O., Trifonov, V. A., Leonard, J. A., Vorobieva, N. V., Ovodov, N. D., Graphodatsky, A. S., and Wayne, R. K. (2013). Ancient DNA analysis affirms the canid from Altai as a primitive dog. PLoS One 8(3), e57754.
| Crossref | Google Scholar | PubMed |
Fan, Z., Silva, P., Gronau, I., Wang, S., Armero, A. S., Schweizer, R. M., Ramirez, O., Pollinger, J., Galaverni, M., Ortega Del-Vecchyo, D., Du, L., Zhang, W., Zhang, Z., Xing, J., Vilà, C., Marques-Bonet, T., Godinho, R., Yue, B., and Wayne, R. K. (2016). Worldwide patterns of genomic variation and admixture in gray wolves. Genome Research 26, 163-73.
| Crossref | Google Scholar | PubMed |
Feuerborn, T. R., Carmagnini, A., Losey, R. J., Nomokonova, T., Askeyev, A., Askeyev, I., Askeyev, O., Antipina, E. E., Appelt, M., Bachura, O. P., Beglane, F., Bradley, D. G., Daly, K. G., Gopalakrishnan, S., Murphy Gregersen, K., Guo, C., Gusev, A. V., Jones, C., Kosintsev, P. A., Kuzmin, Y. V., Mattiangeli, V., Perri, A. R., Plekhanov, A. V., Ramos-Madrigal, J., Schmidt, A. L., Shaymuratova, D., Smith, O., Yavorskaya, L. V., Zhang, G., Willerslev, E., Meldgaard, M., Gilbert, M. T. P., Larson, G., Dalén, L., Hansen, A. J., Sinding, M.-H. S., and Frantz, L. (2021). Modern Siberian dog ancestry was shaped by several thousand years of Eurasian-wide trade and human dispersal. Proceedings of the National Academy of Sciences of the United States of America 118(39), e2100338118.
| Crossref | Google Scholar | PubMed |
Field, M. A., Yadav, S., Dudchenko, O., Esvaran, M., Rosen, B. D., Skvortsova, K., Edwards, R. J., Keilwagen, J., Cochran, B. J., Manandhar, B., Bustamante, S., Rasmussen, J. A., Melvin, R. G., Chernoff, B., Omer, A., Colaric, Z., Chan, E. K. F., Minoche, A. E., Smith, T. P. L., Gilbert, M. T. P., Bogdanovic, O., Zammit, R. A., Thomas, T., Aiden, E. L., and Ballard, J. W. O. (2022). The Australian dingo is an early offshoot of modern breed dogs. Science Advances 8(16), eabm5944.
| Crossref | Google Scholar | PubMed |
Fillios, M. A., and Taçon, P. S. C. (2016). Who let the dogs in? A review of the recent genetic evidence for the introduction of the dingo to Australia and implications for the movement of people. Journal of Archaeological Science: Reports 7, 782-792.
| Crossref | Google Scholar |
Fogg, B. R., Howe, N., and Pierotti, R. (2015). Relationships between Indigenous American Peoples and Wolves 1: Wolves as Teachers and Guides. Journal of Ethnobiology 35(2), 262-285.
| Crossref | Google Scholar |
Frantz, L. A. F., Mullin, V. E., Pionnier-Capitan, M., Lebrasseur, O., Ollivier, M., Perri, A., Linderholm, A., Mattiangeli, V., Teasdale, M. D., Dimopoulos, E. A., Tresset, A., Duffraisse, M., McCormick, F., Bartosiewicz, L., Gál, E., Nyerges, É. A., Sablin, M. V., Bréhard, S., Mashkour, M., Bălăşescu, A., Gillet, B., Hughes, S., Chassaing, O., Hitte, C., Vigne, J.-D., Dobney, K., Hänni, C., Bradley, D. G., and Larson, G. (2016). Genomic and archaeological evidence suggest a dual origin of domestic dogs. Science 352(6290), 1228-1231.
| Crossref | Google Scholar | PubMed |
Freedman, A. H., and Wayne, R. K. (2017). Deciphering the Origin of Dogs: From Fossils to Genomes. Annual Review of Animal Biosciences 5, 281-307.
| Crossref | Google Scholar | PubMed |
Freedman, A. H., Gronau, I., Schweizer, R. M., Ortega-Del Vecchyo, D., Han, E., Silva, P. M., Galaverni, M., Fan, Z., Marx, P., Lorente-Galdos, B., Beale, H., Ramirez, O., Hormozdiari, F., Alkan, C., Vilà, C., Squire, K., Geffen, E., Kusak, J., Boyko, A. R., Parker, H. G., Lee, C., Tadigotla, V., Siepel, A., Bustamante, C. D., Harkins, T. T., Nelson, S. F., Ostrander, E. A., Marques-Bonet, T., Wayne, R. K., and Novembre, J. (2014). Genome Sequencing Highlights the Dynamic Early History of Dogs. PLoS Genetics 10(1), e1004016.
| Crossref | Google Scholar | PubMed |
Freedman, A. H., Schweizer, R. M., Ortega-Del Vecchyo, D., Han, E., Davis, B. W., Gronau, I., Silva, P. M., Galaverni, M., Fan, Z., Marx, P., Lorente-Galdos, B., Ramirez, O., Hormozdiari, F., Alkan, C., Vilà, C., Squire, K., Geffen, E., Kusak, J., Boyko, A. R., Parker, H. G., Lee, C., Tadigotla, V., Siepel, A., Bustamante, C. D., Harkins, T. T., Nelson, S. F., Marques-Bonet, T., Ostrander, E. A., Wayne, R. K., and Novembre, J. (2016). Demographically-Based Evaluation of Genomic Regions under Selection in Domestic Dogs. PLoS Genetics 12(3), e1005851.
| Crossref | Google Scholar | PubMed |
Galibert, F., Quignon, P., Hitte, C., and André, C. (2011). Toward understanding dog evolutionary and domestication history. Comptes Rendus Biologies 334(3), 190-196.
| Crossref | Google Scholar | PubMed |
Gaunitz, C., Fages, A., Hanghøj, K., Albrechtsen, A., Khan, N., Schubert, M., Seguin-Orlando, A., Owens, I. J., Felkel, S., Bignon-Lau, O., De Barros Damgaard, P., Mittnik, A., Mohaseb, A. F., Davoudi, H., Alquraishi, S., Alfarhan, A. H., Al-Rasheid, K., Crubézy, E., Benecke, N., Olsen, S., Brown, D., Anthony, D., Massy, K., Pitulko, V., Kasparov, A., Brem, G., Hofreiter, M., Mukhtarova, G., Baimukhanov, N., Lõugas, L., Onar, V., Stockhammer, P. W., Krause, J., Boldgiv, B., Undrakhbold, S., Erdenebaatar, D., Lepetz, S., Mashkour, M., Ludwig, A., Wallner, B., Merz, V., Merz, I., Zaibert, V., Willerslev, E., Librado, P., Outram, A. K., and Orlando, L. (2018). Ancient genomes revisit the ancestry of domestic and Przewalski’s horses. Science 360(6384), 111-114.
| Crossref | Google Scholar | PubMed |
Gentry, A., Clutton-Brock, J., and Groves, C. P. (2004). The naming of wild animal species and their domestic derivatives. Journal of Archaeological Science 31(5), 645-651.
| Crossref | Google Scholar |
Gojobori, J., Arakawa, N., Xiaokaiti, X., Matsumoto, Y., Matsumura, S., Hongo, H., Ishiguro, N., and Terai, Y. (2024). Japanese wolves are most closely related to dogs and share DNA with East Eurasian dogs. Nature Communications 15(1), 1680.
| Crossref | Google Scholar | PubMed |
Goolmeer, T., Skroblin, A., Grant, C., van Leeuwen, S., Archer, R., Gore-Birch, C., and Wintle, B. A (2022). Recognizing culturally significant species and Indigenous-led management is key to meeting international biodiversity obligations. Conservation Letters 15, e12899.
| Crossref | Google Scholar |
Gopalakrishnan, S., Sinding, M.-H. S., Ramos-Madrigal, J., Niemann, J., Samaniego Castruita, J. A., Vieira, F. G., Carøe, C., Montero, M. d. M., Kuderna, L., Serres, A., González-Basallote, V. M., Liu, Y.-H., Wang, G.-D., Marques-Bonet, T., Mirarab, S., Fernandes, C., Gaubert, P., Koepfli, K.-P., Budd, J., Rueness, E. K., Sillero, C., Heide-Jørgensen, M. P., Petersen, B., Sicheritz-Ponten, T., Bachmann, L., Wiig, Ø., Hansen, A. J., and Gilbert, M. T. P. (2018). Interspecific Gene Flow Shaped the Evolution of the Genus Canis. Current Biology 28(21), 3441-3449.e5.
| Crossref | Google Scholar |
Greig, K., Gosling, A., Collins, C. J., Boocock, J., McDonald, K., Addison, D. J., Allen, M. S., David, B., Gibbs, M., Higham, C. F. W., Liu, F., McNiven, I. J., O’Connor, S., Tsang, C. H., Walter, R., and Matisoo-Smith, E. (2018). Complex history of dog (Canis familiaris) origins and translocations in the Pacific revealed by ancient mitogenomes. Scientific Reports 8(1), 9130.
| Crossref | Google Scholar | PubMed |
Heinsohn, T. (2014). Animal translocation:long-term human influences on the vertebrate zoogeography of Australasia (natural dispersal versus ethnophoresy). Australian Zoologist 32(3), 351-376.
| Crossref | Google Scholar |
Hofman, C. A., and Rick, T. C. (2018). Ancient Biological Invasions and Island Ecosystems: Tracking Translocations of Wild Plants and Animals. Journal of Archaeological Research 26(1), 65-115.
| Crossref | Google Scholar |
ICZN (1999). International Code of Zoological Nomenclature, 4th edn. (International Trust for Zoological Nomenclature: London, U.K.) Available at http://www.iczn.org/iczn/index
Jackson, S. M., Groves, C. P., Fleming, P. J. S., Aplin, K. P., Eldridge, M. D. B., Gonzalez, A., and Helgen, K. M. (2017). The wayward dog: is the Australian native dog or dingo a distinct species? Zootaxa 4317(2), 201-224.
| Crossref | Google Scholar |
Jackson, S. M., Fleming, P. J. S., Eldridge, M. D. B., Ingleby, S., Flannery, T., Johnson, R. N., Cooper, S. J. B., Mitchell, K. J., Souilmi, Y., Cooper, A., Wilson, D. E., and Helgen, K. M. (2019). The Dogma of Dingoes-Taxonomic status of the dingo: A reply to Smith et al. Zootaxa 4564(1), 198-212.
| Crossref | Google Scholar | PubMed |
Jackson, S. M., Fleming, P. J. S., Eldridge, M. D. B., Archer, M., Ingleby, S., Johnson, R. N., and Helgen, K. M. (2020). Taxonomy of the Dingo: It’s an ancient dog. Australian Zoologist 41, 347-357.
| Crossref | Google Scholar |
Koungoulos, L. (2021). Domestication Through Dingo Eyes: An Australian Perspective on Human-Canid Interactions Leading to the Earliest Dogs. Human Ecology 49, 691-705.
| Crossref | Google Scholar |
Koungoulos, L., and Fillios, M. (2020). Hunting dogs down under? On the Aboriginal use of tame dingoes in dietary game acquisition and its relevance to Australian prehistory. Journal of Anthropological Archaeology 58, 101146.
| Google Scholar |
Koungoulos, L. G., Hulme-Beaman, A., Fillios, M., and Willandra Lakes Region World Heritage Aboriginal Advisory Group (2024). Phenotypic diversity in early Australian dingoes revealed by traditional and 3D geometric morphometric analysis. Scientific Reports 14(1),.
| Crossref | Google Scholar |
Kreplins, T. L., Stephens, D., Laing, N., and Oskam, C. (2024). An assessment of dingo ancestry in camp dogs in Western Australia. Australian Journal of Zoology 72(3), ZO24008.
| Crossref | Google Scholar |
Krofel, M., Hatlauf, J., Bogdanowicz, W., Campbell, L. A. D., Godinho, R., Jhala, Y. V., Kitchener, A. C., Koepfli, K. P., Moehlman, P., Senn, H., Sillero-Zubiri, C., Viranta, S., Werhahn, G., and Alvares, F. (2022). Towards resolving taxonomic uncertainties in wolf, dog and jackal lineages of Africa, Eurasia and Australasia. Journal of Zoology 316(3), 155-168.
| Crossref | Google Scholar |
Kumar, M., Conroy, G., Ogbourne, S., Cairns, K., Borburgh, L., and Subramanian, S. (2023). Genomic signatures of bottleneck and founder effects in dingoes. Ecology and Evolution 13(9), e10525.
| Crossref | Google Scholar | PubMed |
Larson, G., and Burger, J. (2013). A population genetics view of animal domestication. Trends in Genetics 29(4), 197-205.
| Crossref | Google Scholar | PubMed |
Letnic, M., Ritchie, E. G., and Dickman, C. R. (2012). Top predators as biodiversity regulators: the dingo Canis lupus dingo as a case study. Biological Reviews 87(2), 390-413.
| Crossref | Google Scholar | PubMed |
Lord, K., Feinstein, M., Smith, B., and Coppinger, R. (2013). Variation in reproductive traits of members of the genus Canis with special attention to the domestic dog (Canis familiaris). Behavioural Processes 92(0), 131-142.
| Crossref | Google Scholar | PubMed |
Lord, K. A., Larson, G., Allaby, R. G., and Karlsson, E. K. (2025). A universally applicable definition for domestication. Proceedings of the National Academy of Sciences of the United States of America 122(22), e2413207122.
| Crossref | Google Scholar | PubMed |
Manwell, C., and Baker, C. M. A. (1984). Domestication of the dog: hunter, food, bed-warmer, or emotional object? Zeitschrift für Tierzüchtung und Züchtungsbiologie 101(1–5), 241-256.
| Crossref | Google Scholar |
Marshall, F. B., Dobney, K., Denham, T., and Capriles, J. M. (2014). Evaluating the roles of directed breeding and gene flow in animal domestication. Proceedings of the National Academy of Sciences of the United States of America 111(17), 6153-6158.
| Crossref | Google Scholar | PubMed |
Martins, R. F., Schmidt, A., Lenz, D., Wilting, A., and Fickel, J. (2018). Human‐mediated introduction of introgressed deer across Wallace’s line: Historical biogeography of Rusa unicolor and R. timorensis. Ecology and Evolution 8(3), 1465-1479.
| Crossref | Google Scholar | PubMed |
National Academies of Sciences, Engineering, and Medicine (2019). ‘Evaluating the Taxonomic Status of the Mexican Gray Wolf and the Red Wolf.’ (The National Academies Press: Washington, DC.) 10.17226/25351
Oetjens, M. T., Martin, A., Veeramah, K. R., and Kidd, J. M. (2018). Analysis of the canid Y-chromosome phylogeny using short-read sequencing data reveals the presence of distinct haplogroups among Neolithic European dogs. BMC Genomics 19(1), 350.
| Crossref | Google Scholar | PubMed |
Oskarsson, M. C. R., Klütsch, C. F. C., Boonyaprakob, U., Wilton, A., Tanabe, Y., and Savolainen, P. (2012). Mitochondrial DNA data indicate an introduction through Mainland Southeast Asia for Australian dingoes and Polynesian domestic dogs. Proceedings of the Royal Society B: Biological Sciences 279(1730), 967-974.
| Crossref | Google Scholar | PubMed |
Parker, H. G., Dreger, D. L., Rimbault, M., Davis, B. W., Mullen, A. B., Carpintero-Ramirez, G., and Ostrander, E. A. (2017). Genomic Analyses Reveal the Influence of Geographic Origin, Migration, and Hybridization on Modern Dog Breed Development. Cell Reports 19(4), 697-708.
| Crossref | Google Scholar | PubMed |
Pendleton, A. L., Shen, F., Taravella, A. M., Emery, S., Veeramah, K. R., Boyko, A. R., and Kidd, J. M. (2018). Comparison of village dog and wolf genomes highlights the role of the neural crest in dog domestication. BMC Biology 16(1), 64.
| Crossref | Google Scholar | PubMed |
Perri, A. R., Feuerborn, T. R., Frantz, L. A. F., Larson, G., Malhi, R. S., Meltzer, D. J., and Witt, K. E. (2021). Dog domestication and the dual dispersal of people and dogs into the Americas. Proceedings of the National Academy of Sciences of the United States of America 118(6), e2010083118.
| Crossref | Google Scholar | PubMed |
Pilot, M. (2021). Disentangling the admixed trails of grey wolf evolution. Molecular Ecology 30(24), 6509-6512.
| Crossref | Google Scholar | PubMed |
Pilot, M., Malewski, T., Moura, A. E., Grzybowski, T., Oleński, K., Ruść, A., Kamiński, S., Fadel, F. R., Mills, D. S., Alagaili, A. N., Mohammed, O. B., Kłys, G., Okhlopkov, I. M., Suchecka, E., and Bogdanowicz, W. (2015). On the origin of mongrels: evolutionary history of free-breeding dogs in Eurasia. Proceedings of the Royal Society B: Biological Sciences 282(1820), 20152189.
| Crossref | Google Scholar | PubMed |
Purugganan, M. D. (2022). What is domestication? Trends in Ecology & Evolution 37(8), 663-671.
| Crossref | Google Scholar | PubMed |
Ritchie, E. G., Smith, B. P., van Eeden, L. M., and Nimmo, D. G. (2018). Species definitions shape policy. Science 361(6409), 1324.
| Crossref | Google Scholar | PubMed |
Sacks, B. N., Brown, S. K., Stephens, D., Pedersen, N. C., Wu, J.-T., and Berry, O. (2013). Y chromosome analysis of dingoes and Southeast Asian village dogs suggests a Neolithic continental expansion from Southeast Asia followed by multiple Austronesian dispersals. Molecular Biology and Evolution 30, 1103-18.
| Crossref | Google Scholar | PubMed |
Savolainen, P., Leitner, T., Wilton, A. N., Matisoo-Smith, E., and Lundeberg, J. (2004). A detailed picture of the origin of the Australian dingo, obtained from the study of mitochondrial DNA. Proceedings of the National Academy of Sciences 101(33), 12387-12390.
| Crossref | Google Scholar |
Scarsbrook, L., Cairns, K. M., Mitchell, K. J., Bougiouri, K., Evin, A., Harris, A. C., Wood, A. E., Zehui, Zhang, Z., Alves, J. M., Lawson, D. J., Ditchfield, P. W., Granja-Martins, S., Styring, A. K., Tabadda, K., Thalmann, O., Crowther, M. S., Curry, M., Feuerborn, T. R., Koungoulos, L. G., Letnic, M., Reed, E. H., Sabin, R., Parker, H. G., Ostrander, E. A., Frantz, L. A. F., Larson, G., and Fillios, M. A. (In Press). The Impacts of European Arrival on Australian Dingoes. Proceedings of the National Academy of Sciences. Accepted 09/07/2025.
Schweizer, R. M., Durvasula, A., Smith, J., Vohr, S. H., Stahler, D. R., Galaverni, M., Thalmann, O., Smith, D. W., Randi, E., Ostrander, E. A., Green, R. E., Lohmueller, K. E., Novembre, J., and Wayne, R. K. (2018). Natural Selection and Origin of a Melanistic Allele in North American Gray Wolves. Molecular Biology and Evolution 35(5), 1190-1209.
| Crossref | Google Scholar |
Shannon, L. M., Boyko, R. H., Castelhano, M., Corey, E., Hayward, J. J., McLean, C., White, M. E., Abi Said, M., Anita, B. A., Bondjengo, N. I., Calero, J., Galov, A., Hedimbi, M., Imam, B., Khalap, R., Lally, D., Masta, A., Oliveira, K. C., Pérez, L., Randall, J., Tam, N. M., Trujillo-Cornejo, F. J., Valeriano, C., Sutter, N. B., Todhunter, R. J., Bustamante, C. D., and Boyko, A. R. (2015). Genetic structure in village dogs reveals a Central Asian domestication origin. Proceedings of the National Academy of Sciences of the United States of America 112(44), 13639-13644.
| Crossref | Google Scholar | PubMed |
Shipman, P. (2021). What the dingo says about dog domestication. The Anatomical Record 304(1), 19-30.
| Crossref | Google Scholar | PubMed |
Sinding, M.-H. S., Gopalakrishnan, S., Ramos-Madrigal, J., de Manuel, M., Pitulko, V. V., Kuderna, L., Feuerborn, T. R., Frantz, L. A. F., Vieira, F. G., Niemann, J., Samaniego Castruita, J. A., Carøe, C., Andersen-Ranberg, E. U., Jordan, P. D., Pavlova, E. Y., Nikolskiy, P. A., Kasparov, A. K., Ivanova, V. V., Willerslev, E., Skoglund, P., Fredholm, M., Wennerberg, S. E., Heide-Jørgensen, M. P., Dietz, R., Sonne, C., Meldgaard, M., Dalén, L., Larson, G., Petersen, B., Sicheritz-Pontén, T., Bachmann, L., Wiig, Ø., Marques-Bonet, T., Hansen, A. J., and Gilbert, M. T. P. (2020). Arctic-adapted dogs emerged at the Pleistocene–Holocene transition. Science 368(6498), 1495-1499.
| Crossref | Google Scholar | PubMed |
Skoglund, P., Ersmark, E., Palkopoulou, E., and Dalén, L. (2015). Ancient wolf genome reveals an early divergence of domestic dog ancestors and admixture into high-latitude breeds. Current Biology 25(11), 1515-1519.
| Crossref | Google Scholar | PubMed |
Smith, B. P., and Litchfield, C. A. (2009). A Review of the Relationship between Indigenous Australians, Dingoes (Canis dingo) and Domestic Dogs (Canis familiaris). Anthrozoös 22(2), 111-128.
| Google Scholar |
Smith, B. P., Cairns, K. M., Adams, J. W., Newsome, T. M., Fillios, M., Déaux, E. C., Parr, W. C. H., Letnic, M., van Eeden, L. M., Appleby, R. G., Bradshaw, C., Savolainen, P., Ritchie, E. G., Nimmo, D. G., Archer-Lean, C., Greenville, A. C., Dickman, C. R., Watson, L., Moseby, K. E., Doherty, T. S., Wallach, A. D., Morrant, D. S., and Crowther, M. S. (2019). Taxonomic status of the Australian dingo: the case for Canis dingo Meyer, 1793. Zootaxa 4564(1), 173-197.
| Crossref | Google Scholar | PubMed |
Smith, B. P., Appleby, R. G., and Cairns, K. M. (2023). Reimagining the Dingo: The ‘Australian Wolf ’ or Just a Feral Dog?. In ‘The Wolf’. (Eds I. Convery, O. Nevin, E. van Maanen, et al) pp. 161-178. (Boydell and Brewer.) 10.1515/9781805430551-020
Souilmi, Y., Wasef, S., Williams, M. P., Conroy, G., Bar, I., Bover, P., Dann, J., Heiniger, H., Llamas, B., Ogbourne, S., Archer, M., Ballard, J. W. O., Reed, E., Tobler, R., Koungoulos, L., Walshe, K., Wright, J. L., Balme, J., O’Connor, S., Cooper, A., and Mitchell, K. J. (2024). Ancient genomes reveal over two thousand years of dingo population structure. Proceedings of the National Academy of Sciences of the United States of America 121(30), e2407584121.
| Crossref | Google Scholar | PubMed |
Stephens, D., Wilton, A. N., Fleming, P. J. S., and Berry, O. (2015). Death by sex in an Australian icon: a continent-wide survey reveals extensive hybridization between dingoes and domestic dogs. Molecular Ecology 24(22), 5643-5656.
| Crossref | Google Scholar | PubMed |
Stephens, D., Fleming, P. J. S., Sawyers, E., and Mayr, T. P. (2022). An isolated population reveals greater genetic structuring of the Australian dingo. Scientific Reports 12(1), 19105.
| Crossref | Google Scholar | PubMed |
Surbakti, S., Parker, H. G., McIntyre, J. K., Maury, H. K., Cairns, K. M., Selvig, M., Pangau-Adam, M., Safonpo, A., Numberi, L., Runtuboi, D. Y. P., Davis, B. W., and Ostrander, E. A. (2020). New Guinea highland wild dogs are the original New Guinea singing dogs. Proceedings of the National Academy of Sciences of the United States of America 117(39), 24369-24376.
| Crossref | Google Scholar | PubMed |
Thalmann, O., Shapiro, B., Cui, P., Schuenemann, V. J., Sawyer, S. K., Greenfield, D. L., Germonpré, M. B., Sablin, M. V., López-Giráldez, F., Domingo-Roura, X., Napierala, H., Uerpmann, H.-P., Loponte, D. M., Acosta, A. A., Giemsch, L., Schmitz, R. W., Worthington, B., Buikstra, J. E., Druzhkova, A., Graphodatsky, A. S., Ovodov, N. D., Wahlberg, N., Freedman, A. H., Schweizer, R. M., Koepfli, K.-P., Leonard, J. A., Meyer, M., Krause, J., Pääbo, S., Green, R. E., and Wayne, R. K. (2013). Complete mitochondrial genomes of ancient canids suggest a European origin of domestic dogs. Science 342(6160), 871-874.
| Crossref | Google Scholar | PubMed |
Thiele, K. R., Conix, S., Pyle, R. L., Barik, S. K., Christidis, L., Costello, M. J., van Dijk, P. P., Kirk, P., Lien, A., Thomson, S. A., Zachos, F. E., Zhang, Z.-Q., and Garnett, S. T. (2021). Towards a global list of accepted species I. Why taxonomists sometimes disagree, and why this matters. Organisms Diversity & Evolution 21(4), 615-622.
| Crossref | Google Scholar |
Thomson, P. C., Rose, K., and Kok, N. E. (1992). The behavioural ecology of dingoes in north-Western Australia. V. Population dynamics and variation in the social system. Wildlife Research 19(5), 565-583.
| Google Scholar |
Veron, G., Willsch, M., Dacosta, V., Patou, M.-L., Seymour, A., Bonillo, C., Couloux, A., Wong, S. T., Jennings, A. P., Fickel, J., and Wilting, A. (2014). The distribution of the Malay civet Viverra tangalunga (Carnivora: Viverridae) across Southeast Asia: natural or human-mediated dispersal? Zoological Journal of the Linnean Society 170(4), 917-932.
| Crossref | Google Scholar |
vonHoldt, B. M., and Aardema, M. L. (2020). Updating the bibliography of interbreeding among Canis in North America. Journal of Heredity 111, 249-262.
| Crossref | Google Scholar | PubMed |
vonHoldt, B. M., Pollinger, J. P., Lohmueller, K. E., Han, E., Parker, H. G., Quignon, P., Degenhardt, J. D., Boyko, A. R., Earl, D. A., Auton, A., Reynolds, A., Bryc, K., Brisbin, A., Knowles, J. C., Mosher, D. S., Spady, T. C., Elkahloun, A., Geffen, E., Pilot, M., Jedrzejewski, W., Greco, C., Randi, E., Bannasch, D., Wilton, A., Shearman, J., Musiani, M., Cargill, M., Jones, P. G., Qian, Z., Huang, W., Ding, Z.-L., Zhang, Y.-p., Bustamante, C. D., Ostrander, E. A., Novembre, J., and Wayne, R. K. (2010). Genome-wide SNP and haplotype analyses reveal a rich history underlying dog domestication. Nature 464, 898-902.
| Crossref | Google Scholar | PubMed |
vonHoldt, B., Pollinger, J., Earl, D., Parker, H., Ostrander, E., and Wayne, R. (2013). Identification of recent hybridization between gray wolves and domesticated dogs by SNP genotyping. Mammalian Genome 24(1–2), 80-88.
| Crossref | Google Scholar | PubMed |
vonHoldt, B. M., Brzeski, K. E., Wilcove, D. S., and Rutledge, L. Y. (2018). Redefining the role of admixture and genomics in species conservation. Conservation Letters 11(2), e12371.
| Crossref | Google Scholar |
Weeks, A. R., Kriesner, P., Bartonicek, N., van Rooyen, A., Cairns, K. M., and Ahrens, C. W. (2024). Genetic structure and common ancestry expose the dingo-dog hybrid myth. Evolution Letters 9, 1-12.
| Crossref | Google Scholar | PubMed |
Zeder, M. A. (2012). The Domestication of Animals. Journal of Anthropological Research 68(2), 161-190.
| Crossref | Google Scholar |
Zeller, U., and Göttert, T. (2019). The relations between evolution and domestication reconsidered - Implications for systematics, ecology, and nature conservation. Global Ecology and Conservation 20, e00756.
| Crossref | Google Scholar |
Zhang, S.-j., Wang, G.-D., Ma, P., Zhang, L.-l., Yin, T.-T., Liu, Y.-h., Otecko, N. O., Wang, M., Ma, Y.-p., Wang, L., Mao, B., Savolainen, P., and Zhang, Y.-p. (2020). Genomic regions under selection in the feralization of the dingoes. Nature Communications 11(1), 671.
| Crossref | Google Scholar | PubMed |


