114 SPECIFIC AMINO ACIDS INFLUENCE LINEAGE DIFFERENTIATION IN MOUSE BLASTOCYSTS AND OUTGROWTHS
R. Sihota A , S. Brooks A , C. Tong A , I. T. Cameron B , T. P. Fleming A and J. J. Eckert A BA School of Biological Sciences, University of Southampton, Southampton, UK;
B DOHaD School of Medicine, University of Southampton, Southampton, UK
Reproduction, Fertility and Development 20(1) 137-137 https://doi.org/10.1071/RDv20n1Ab114
Published: 12 December 2007
Abstract
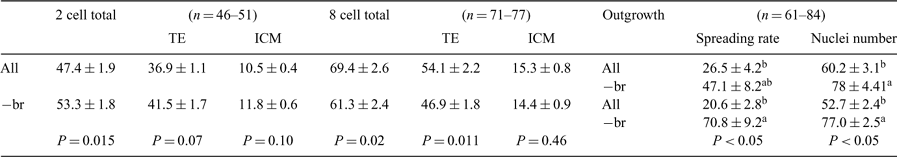
Blastocyst biogenesis and cell lineage differentiation into inner cell mass and trophectoderm (TE) are highly regulated processes initiated during cleavage. Cell positioning and contact patterns are well-accepted broad extrinsic factors involved in regulating blastocyst formation; however, subtle changes in the chemical composition of culture media, similarly extrinsic, may exert significant consequences for differentiation. In this study, we have examined whether, when, and how the lack of branched chain amino acids impacts on blastocyst differentiation and TE development. Using superovulated MF1 mice, embryos were flushed at 50 (2 cell) or 75 (8 cell) h after hCG and cultured in mKSOM with 0.6% BSA and amino acids at uterine fluid concentrations (Porter et al. 2003 Pediatr. Res. 53, 46A) with (all) or without (–br) the branched-chain amino acids valine, leucine, and isoleucine. Blastocysts (60 or 36 h after culture onset) were differentially labeled using the TNBS-anti-DNP-complement method with propidium iodide and Hoechst, and cells were counted on z-series with overlays using Metamorph software. For outgrowths, blastocysts developed with all aa or –br from 8 cells for 36 h were placed into KSOM supplemented with 10% FCS and either all aa or –br and cultured for an additional 120 h, scored daily, and 4′,6-diamidino-2-phenylindole nuclei counts were made at 120 h. Lack of branched amino acids from the 2-cell stage onwards caused a significant increase in total blastocyst cells with predominantly higher TE numbers, whereas, when exposed to –br from the 8 cell stage onwards, the outcome was reversed (ANOVA; Table 1). Upon outgrowth, prior amino acid composition was relatively less influential in determining spreading pattern and rate over time compared to outgrowth amino acid conditions (ANOVA; Table 1). Our data indicate that lineage development in the blastocyst is sensitive to small changes in free amino acids available and responds differently according to exposure time and duration. Subsequent development predominantly depends upon outgrowth conditions, irrespective of earlier experiences, suggesting that a similar capability for adaptive changes in response to different outgrowth media is maintained. These findings may help understand developmental plasticity and consider dynamic and potential longer-term responses to minor culture changes.
|
Funding by DOHaD, Gerald Kerkut Trust, and NIH are gratefully acknowledged.


