31 PRODUCTION EFFICIENCY OF GENE KNOCKOUT PIGS USING GENOME EDITING AND SOMATIC CELL CLONING
H. Matsunari A B , M. Watanabe A B , K. Nakano B B , A. Uchikura B , Y. Asano B , S. Hatae B , T. Takeishi B , K. Umeyama A B , M. Nagaya A B , S. Miyagawa D , Y. Hanazono E , H. Nakauchi F and H. Nagashima A BA Meiji University International Institute for Bio-Resource Research (MUIIBR), Kawasaki, Japan;
B Laboratory of Developmental Engineering, Department of Life Sciences, School of Agriculture, Meiji University, Kawasaki, Japan;
C JSPS Research Fellow, Tokyo, Japan;
D Division of Organ Transplantation, Department of Surgery, Osaka University Graduate School of Medicine, Osaka, Japan;
E Division of Regenerative Medicine, Center for Molecular Medicine, Jichi Medical University, Tochigi, Japan;
F Center for Stem Cell and Regenerative Medicine, Institute of Medical Science, The University of Tokyo, Tokyo, Japan
Reproduction, Fertility and Development 27(1) 108-108 https://doi.org/10.1071/RDv27n1Ab31
Published: 4 December 2014
Abstract
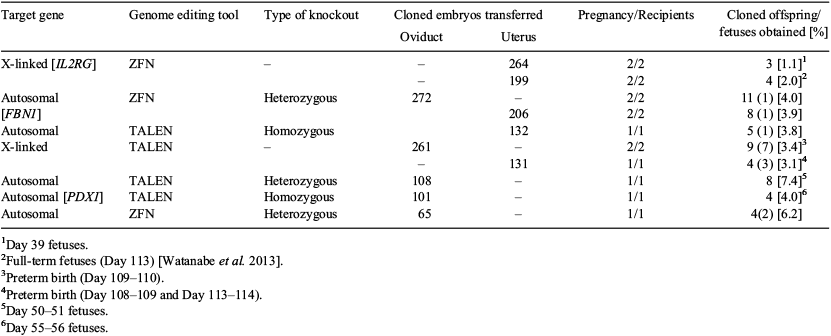
Genome editing technologies have been used as a powerful strategy for the generation of genetically modified pigs. We previously developed genetically modified clone pigs with organogenesis-disabled phenotypes, as well as pigs exhibiting diseases with similar features to those of humans. Here, we report the production efficiency of various gene knockout cloned pigs from somatic cells that were genetically modified using zinc finger nucleases (ZFN) or transcription activator-like effector nucleases (TALEN). The ZFN- or TALEN-encoding mRNAs, which targeted 7 autosomal or X-linked genes, were introduced into porcine fetal fibroblast cells using electroporation. Clonal cell populations carrying induced mutations were selected after limiting dilution. The targeted portion of the genes was amplified using PCR, followed by sequencing and mutation analysis. Among the collected knockout cell colonies, cells showing good proliferation and morphology were selected and used for somatic cell nuclear transfer (SCNT). In vitro-matured oocytes were obtained from porcine cumulus-oocyte complexes cultured in NCSU23-based medium and were used to obtain recipient oocytes for SCNT after enucleation. SCNT was performed as reported previously (Matsunari et al. 2008). The cloned embryos were cultured for 7 days in porcine zygote medium (PZM)-5 to assess their developmental ability. Cloned embryos were transplanted into the oviduct or uterus of oestrus-synchronized recipient gilts to evaluate their competence to develop to fetuses or piglets. Cloned embryos reconstructed with 7 types of knockout cells showed equal development to blastocysts compared with those derived from the wild-type cells (54.5–83.3% v. 60.7%). Our data (Table 1) demonstrated that the reconstructed embryos derived from knockout cells could efficiently give rise to cloned offspring regardless of the type of genome editing methodology (i.e. ZFN or TALEN).
|
This study was supported by JST, ERATO, the Nakauchi Stem Cell and Organ Regeneration Project, JST, CREST, Meiji University International Institute for Bio-Resource Research (MUIIBR), and JSPS KAKENHI Grant Number 26870630.


